
Fairspace is a secure place for managing research data.
Research teams have their own workspaces in which they
can manage research data collections.
Researchers can upload directories and files to data collections.
Data access is organised on data collection level.
Collections can be shared with other teams or individual researchers.
Also, collections can be published for all researchers in the organisation.
Collections and files can be annotated with descriptive metadata. The metadata is stored using the Resource Description Framework (RDF) in an Apache Jena database. For the metadata, a data model can be configured that suits the data management needs of the organisation. The data model is specified using the Shapes Constraint Language (SHACL), see the section on Data model and view configuration. Descriptive metadata entities (e.g., subjects, projects, samples) should be added to the database by a careful process, ensuring that duplicates and inconsistencies are avoided and that all entities have proper unique identifiers. The application provides overviews of the available metadata entities. In the collection browser, researchers can link their collections and files to these entities or add textual descriptions and key words.
Usage
User interface
Login
Users are authenticated using Keycloak, an open-source identity provider that provides secure authentication methods and can be configured to integrate with institutional identity providers using user federation or identity brokering, see the Keycloak server administration pages.
The user either logs in directly using Keycloak or is forwarded to a configured external login:
Workspaces
Users enter Fairspace on the workspaces page that lists all workspaces. A workspace represents a team in the organisation that collaborates on research data collections.
Workspace administrators can edit the workspace overview page and manage workspace membership. All workspace members can add collections to the workspace.
Collections
The contents of collections can be navigated in the collections browser. It behaves like a regular file browser. Click to select a directory or file and see its metadata, double click to navigate into directories or open a file.
Access is managed on collection level. Users with at least write access to a collection can upload files or directories, rename or delete files, restore old file versions, and edit the associated metadata.
Users with manage access can share collections with other users or workspaces, and change the default access mode for workspace members. Collection managers can also change the status of the collection (Active, Read-only or Archived), change the view mode (Restricted, Metadata published or Data published — only available in the Read-only status), delete the collection, or transfer ownership of the collection to another workspace.
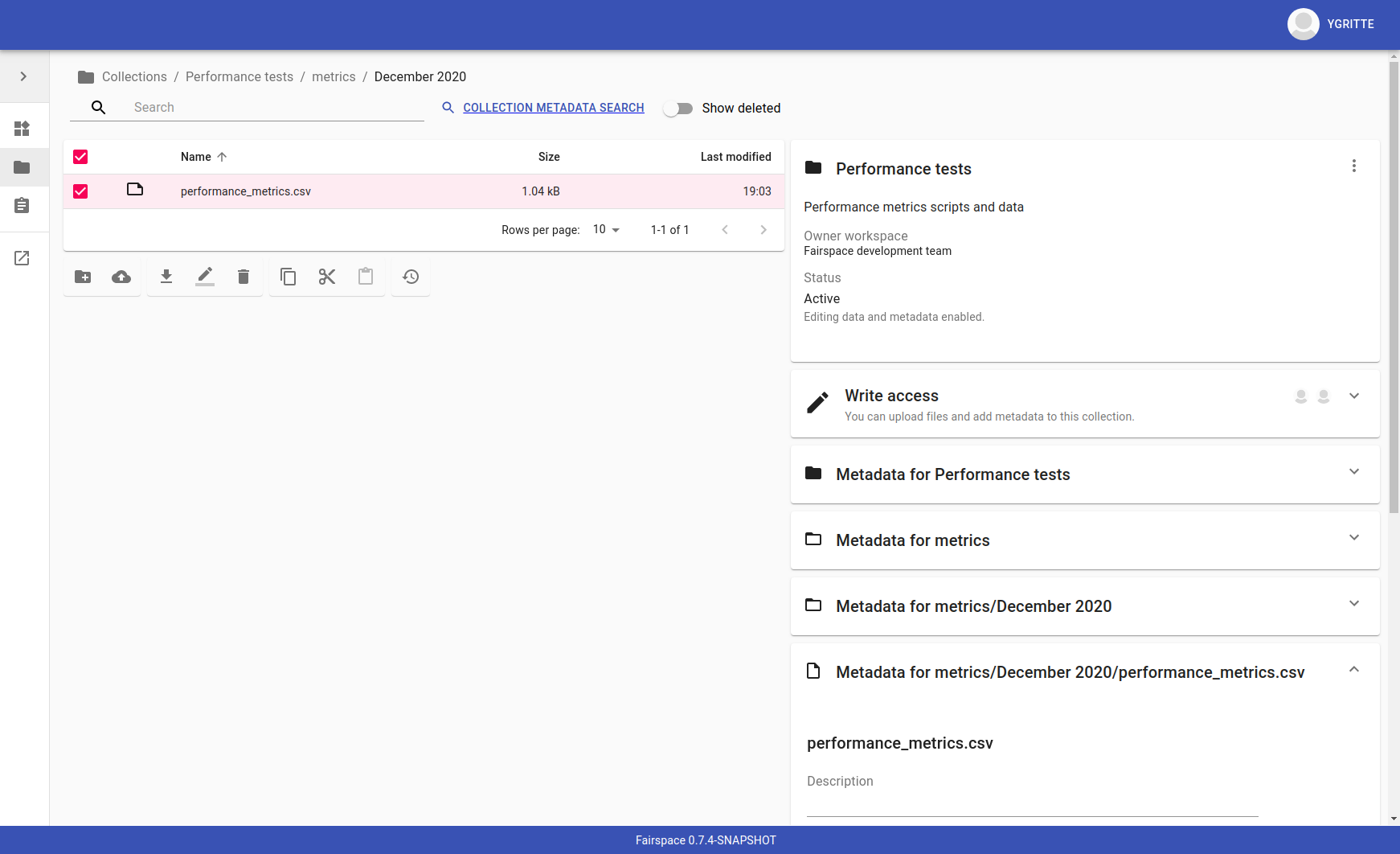
Due to the data loss prevention, data in Fairspace is not removed from the system on deletion. Deleted collections and files can still be viewed in the application using "Show deleted" switch. The goal is to prevent deleted data from being overwritten by users (not to create collections or files with the paths that already existed in the system) and to allow administrators to perform special actions (to be performed only in exceptional special cases), like undeletion or permanent removal, to revert accidental removal or creation of a collection or a file.
Metadata forms
Users with write access to the collection can annotate collections, directories and files using metadata forms. Free text fields, like description and key words, can be entered freely, links to shared entities, like subjects, samples and projects, or values from a controlled vocabulary, like taxonomy or analysis type, can be selected from a list:
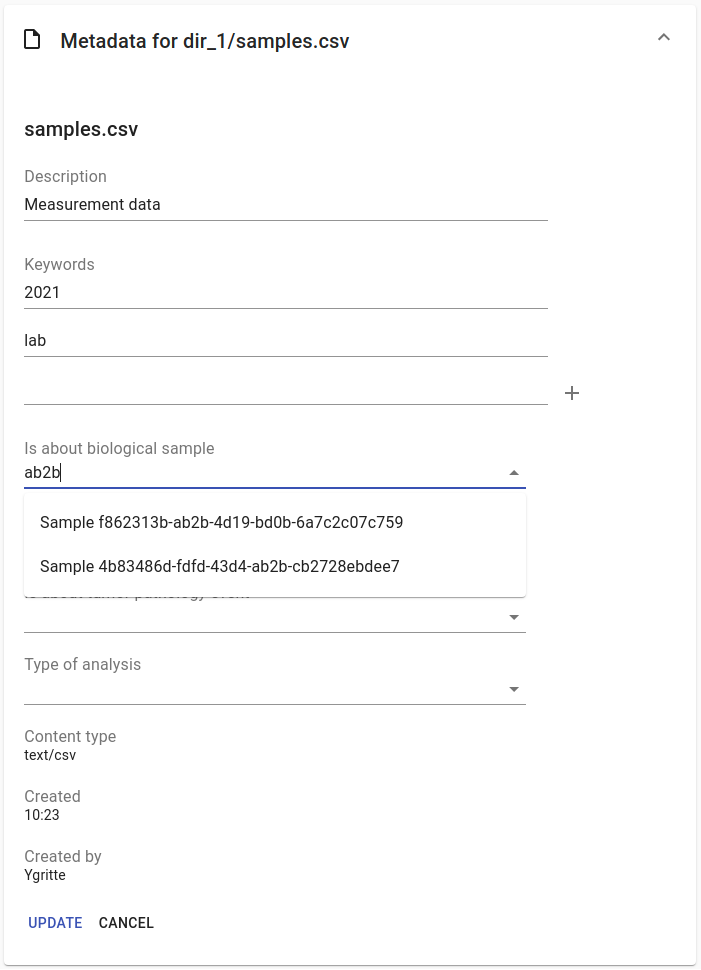
The shared metadata entities and controlled vocabularies cannot be added via the user interface. The RDF metadata API should be used for that instead.
Metadata upload
Another way to annotate directories and files is by uploading a comma-separated values (CSV) file with metadata. This section describes the CSV-based format used for bulk metadata uploads.
The file should be a valid CSV-file:
-
Records are separated with a
,-character. -
Values may be enclosed in double quotes:
"value". -
In values that contain a double, the double quotes need to be escaped by replacing them with double double quotes:
Example "quoted" textbecomes"Example ""quoted"" text".
In the metadata upload, lines starting with # are ignored. These lines are considered to be comments.
The file should have a header row containing the names of the columns.
The mandatory Path column is used for the file path. For the property columns, the name should match exactly the name of the property in the database.
The format of the values is as follows:
-
Path: the relative path to a file or a directory (relative to the collection or directory where the file is uploaded). Use
./for the current directory or collection. -
Entity types can be referenced by ID or unique label.
-
Multiple values must be separated by the pipe symbol
|, e.g., usetest|labto enter the valuestestandlab.
The file can be uploaded to the current directory by dropping the file in the metadata panel of the directory, or by selecting the metadata upload button.
By hovering over the metadata upload button, a link to a metadata template file becomes available:
The file describes the format in commented lines and contains the available properties in the header row.
An example comma-separated values file with metadata about the current directory ./,
which is annotated with a description and two key words (sample and lab),
and the file test.txt which is linked to Subject 1 by the unique subject label
and to the RNA-seq analysis type by the analysis type identifier (O6-12).
Path,Is about subject,Type of analysis,Description,Keywords
./,,,Directory with samples,sample|lab,
test.txt,Subject 1,https://institut-curie.org/analysis#O6-12,,This specifies the table:
| Path | Is about subject | Type of analysis | Description | Keywords |
|---|---|---|---|---|
./ |
Directory with samples |
sample|lab |
||
test.txt |
Subject 1 |
Metadata
Explore metadata and find associated collections and files.
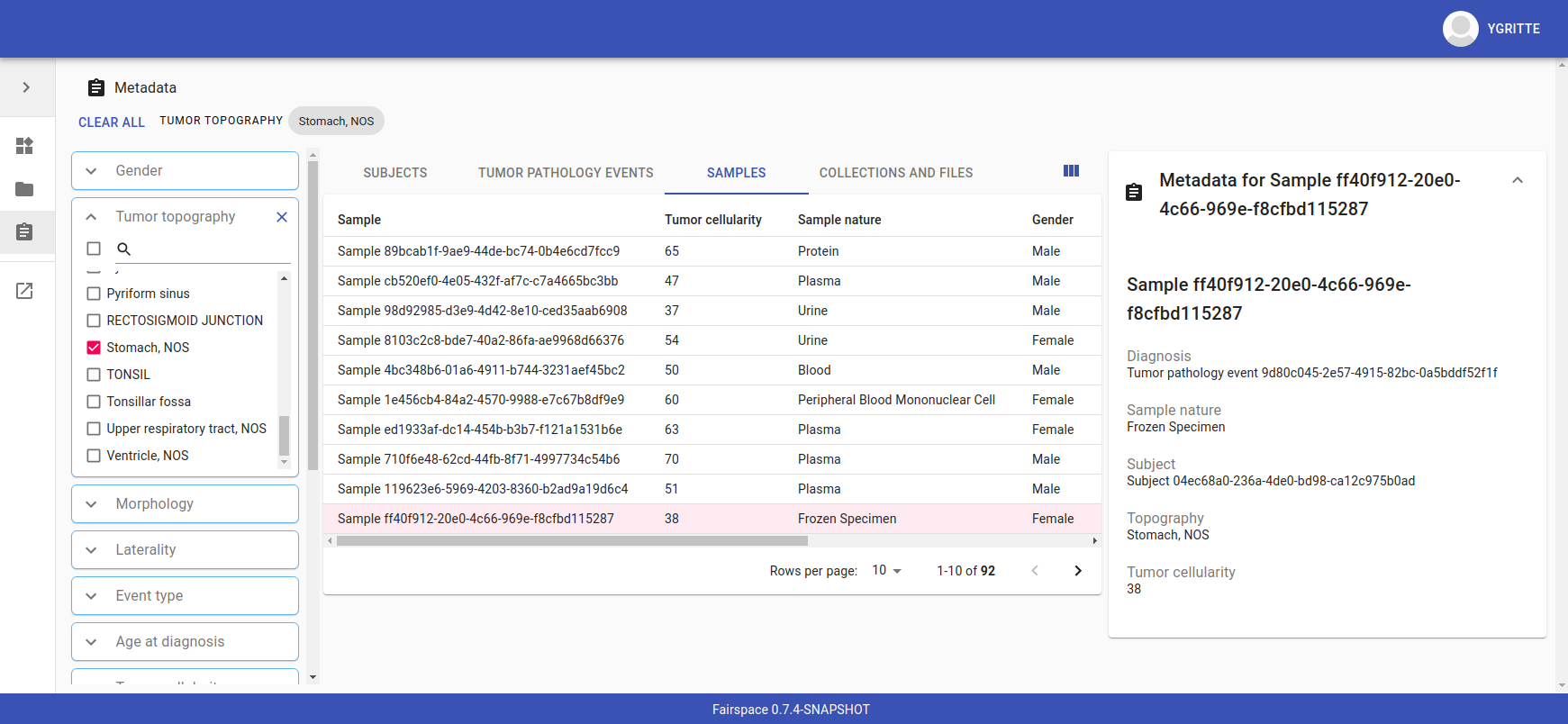
Interfaces for accessing and querying data (API)
The data in Fairspace can be accessed via Application Programming Interfaces (APIs). The user interfaces application uses those APIs, but also other programs can use them, e.g., for automated data uploading or for exporting data for further processing or for synchronisation with other systems.
Authentication
All API endpoints require authentication via an authorisation header. To enable WebDAV clients to connect to Fairspace, also so-called Basic authentication is supported.
For secure authentication, it is strongly advised to use the OpenID Connect (OIDC) / OAuth2 workflow. The user interface application also uses this workflow.
When using the APIs in automated scripts, ensure that an account is used with only the required privileges (conform the principle of least privilege). I.e., when an admin account is not needed, use a non-admin account. For adding shared metadata, an account with Add shared metadata role is required, see Uploading metadata.
When an action is done on behalf of a specific user, do not use a service account or system account for the action directly, but obtain a token for that user first, e.g., by using the impersonation feature of Keycloak. That way the audit logging still captures which user did what.
OpenID Connect (OIDC) / OAuth2 workflow
Fairspace supports OpenID Connect authentication via Keycloak. The workflow for API access is roughly as follows.
-
The client authenticates with the token endpoint of the identity provider (Keycloak) and obtains a signed access token
-
The client uses the access token in the request header when connecting to the Fairspace API
-
Fairspace receives the request with the access token and validates if the token is valid, using the public key of the identity provider.
The token endpoint of Keycloak supports refreshing the token if it is close to expiry. However, checking the token expiration and refreshing make the authentication logic quite complex.
You can either obtain a fresh token before every API request or use an existing library that implements the authentication workflow. For finding available client-side libraries, check the Securing applications and services guide of Keycloak.
For use in scripts, it is advised to obtain a token for offline access, using the Offline access feature of OpenID Connect.
Code to obtain the OpenID Connect authorisation header (Python)
import logging
import os
import requests
log = logging.getLogger()
def fetch_access_token(keycloak_url: str = os.environ.get('KEYCLOAK_URL'),
realm: str = os.environ.get('KEYCLOAK_REALM'),
client_id: str = os.environ.get('KEYCLOAK_CLIENT_ID'),
client_secret: str = os.environ.get('KEYCLOAK_CLIENT_SECRET'),
username: str = os.environ.get('KEYCLOAK_USERNAME'),
password: str = os.environ.get('KEYCLOAK_PASSWORD')) -> str:
"""
Obtain access token from Keycloak
:return: the access token as string.
"""
params = {
'client_id': client_id,
'client_secret': client_secret,
'username': username,
'password': password,
'grant_type': 'password'
}
headers = {
'Content-type': 'application/x-www-form-urlencoded',
'Accept': 'application/json'
}
response = requests.post(f'{keycloak_url}/realms/{realm}/protocol/openid-connect/token',
data=params,
headers=headers)
if not response.ok:
log.error('Error fetching token!', response.json())
raise Exception('Error fetching token.')
data = response.json()
token = data['access_token']
log.info(f"Token obtained successfully. It will expire in {data['expires_in']} seconds")
return token
def auth():
return f'Bearer {fetch_access_token()}'Code to obtain the OpenID Connect authorisation header (bash, curl)
Requires the jq JSON parser.
fetch_access_token() {
curl -s \
--data-urlencode "client_id=${KEYCLOAK_CLIENT_ID}" \
--data-urlencode "client_secret=${KEYCLOAK_CLIENT_SECRET}" \
--data-urlencode "username=${KEYCLOAK_USERNAME}" \
--data-urlencode "password=${KEYCLOAK_PASSWORD}" \
-d 'grant_type=password' \
"${KEYCLOAK_URL}/realms/${KEYCLOAK_REALM}/protocol/openid-connect/token" | jq -r '.access_token'
}
ACCESS_TOKEN=$(fetch_access_token)Basic authentication
For WebDAV client access and for a simpler authentication method
during testing, Fairspace also supports Basic authentication,
which means that the base64 encoded username:password string is sent in the Authorization header together with a prefix Basic .
This authentication method is considered to be less secure than token based authentication, because it requires scripts to have a plain text password stored somewhere. Also, users may have to retype their passwords when logging in, tempting them to choose less secure, easier to remember, passwords.
Code to generate the Basic authorisation header (Python)
import base64
import os
def auth():
username = os.environ.get('KEYCLOAK_USERNAME')
password = os.environ.get('KEYCLOAK_PASSWORD')
return f"Basic {base64.b64encode(f'{username}:{password}'.encode()).decode()}"Code to generate the Basic authorisation header (bash)
AUTH_HEADER="Basic $(echo -n "${KEYCLOAK_USERNAME}:${KEYCLOAK_PASSWORD}" | base64)"Examples
In the examples in this documentation, we assume one of both methods to be available.
This means for the Python examples that a function auth() should be implemented that returns the authorisation header value, see the examples above.
import os
from requests import Response, Session
def auth():
""" Returns authorisation header
Replace this with an implementation from one of the sections above.
"""
pass
server_url = os.environ.get('FAIRSPACE_URL')
headers = {
'Authorization': auth()
}
response = Session().get(f'{server_url}/api/users/current', headers=headers)
if not response.ok:
raise Exception(f"Error fetching current user: {response.status_code} {response.reason}")
print(response.json())For examples using curl, an authorisation header needs to be passed using the -H option.
For Basic authentication:
AUTH_HEADER="Basic $(echo -n "${KEYCLOAK_USERNAME}:${KEYCLOAK_PASSWORD}" | base64)"
curl -i -H "Authorization: ${AUTH_HEADER}" "${FAIRSPACE_URL}/api/users/current"For OpenID Connect:
# ACCESS_TOKEN=...
AUTH_HEADER="Bearer ${ACCESS_TOKEN}"
curl -i -H "Authorization: ${AUTH_HEADER}" "${FAIRSPACE_URL}/api/users/current"Automatic authentication in Jupyter Hub
In Jupyter Hub, users are automatically authenticated and can directly connect to the local API address without adding authentication headers.
WebDAV
A file storage API is exposed via the WebDAV protocol for accessing the file system via the web. It runs on /api/webdav/.
This endpoint can be used by many file explorers,
including Windows Explorer,
and by tools like FileZilla and Cyberduck.
Use https://fairspace.example.com/api/webdav/ or
davs://fairspace.example.com/api/webdav/ as location, with
fairspace.example.com replaced by the server name.
All visible collections in the system are exposed as top-level directories. Creating a top-level directory via WebDAV will result in an error message, see Create collection or directory.
The Web-based Distributed Authoring and Versioning (WebDAV) protocol allows users to operate on collections and files. Fairspace exposes a WebDAV API for accessing the file systems, while restricting access to only the files accessible by the user.
The WebDAV API allows to upload and download files and to perform standard file operations such as copying or moving, as well as custom operations, such as collection lifecycle management and advanced data loss prevention features such as versioning and undeletion.
Be aware that the move operation moves both file content and all its metadata (e.g. linked metadata entities), whereas copy includes only the file content and standard webdav properties, like file size.
Directory listing and path properties
PROPFIND /api/webdav/{path} |
|
|---|---|
Request headers: |
|
|
When |
|
Include deleted paths when the value is |
|
Specify a version number to request properties of a specific file version.
The first version has number |
|
Include list of metadata entities that are linked to the resource, when value |
Request body: |
|
To include also custom Fairspace attributes in the response, like the collection description, send the following request body: |
|
Code examples
Check if path exists (Python)
import logging
import os
from requests import Request, Response, Session
log = logging.getLogger()
server_url = os.environ.get('FAIRSPACE_URL')
def exists(path):
""" Check if a path exists
"""
headers = {
'Depth': '0',
'Authorization': auth()
}
session = Session()
req = Request('PROPFIND', f'{server_url}/api/webdav/{path}/', headers=headers, cookies=session.cookies)
response: Response = session.send(req.prepare())
return response.okFetch directory listing (Python)
import logging
import os
from requests import Request, Response, Session
from xml.etree.ElementTree import fromstring
log = logging.getLogger()
server_url = os.environ.get('FAIRSPACE_URL')
def ls(path: str):
""" List contents of path
"""
headers = {
'Depth': '1',
'Authorization': auth()
}
session = Session()
req = Request('PROPFIND', f'{server_url}/api/webdav/{path}', headers=headers, cookies=session.cookies)
response: Response = session.send(req.prepare())
if not response.ok:
raise Exception(f"Error fetching directory '{path}': {response.status_code} {response.reason}")
tree = fromstring(response.content.decode())
for item in tree.findall('{DAV:}response'):
print(item.find('{DAV:}href').text)Fetch directory listing (curl)
Requires the xmlstarlet tool.
curl -s -H "Authorization: ${AUTH_HEADER}" -X PROPFIND -H "Depth: 1" "${FAIRSPACE_URL}/api/webdav/${path}" -d '<propfind><allprop /></propfind>' \
| xmlstarlet sel -T -t -m d:multistatus/d:response -v d:propstat/d:prop/d:displayname -nExample response
Example PROPFIND response
Example response using PROPFIND on the root location https://fairspace.ci.fairway.app/api/webdav with Depth: 1 and request body <propfind><allprop /></propfind>.
Adding the <allprop /> in the request results in custom Fairspace properties,
like the description (ns1:comment), to be included in the WebDAV response.
<?xml version="1.0" encoding="utf-8" ?>
<d:multistatus xmlns:ns1="https://fairspace.nl/ontology#" xmlns:d="DAV:">
<d:response>
<d:href>/api/webdav/</d:href>
<d:propstat>
<d:prop>
<d:getcontenttype></d:getcontenttype>
<d:getetag>"https://fairspace.ci.fairway.app/api/webdav"</d:getetag>
<d:iscollection>TRUE</d:iscollection>
<d:displayname></d:displayname>
<d:isreadonly>TRUE</d:isreadonly>
<d:name></d:name>
<d:supported-report-set></d:supported-report-set>
<d:resourcetype>
<d:collection/>
</d:resourcetype>
</d:prop>
<d:status>HTTP/1.1 200 OK</d:status>
</d:propstat>
</d:response>
<d:response>
<d:href>/api/webdav/Demonstration/</d:href>
<d:propstat>
<d:prop>
<ns1:access>Write</ns1:access>
<ns1:canRead>TRUE</ns1:canRead>
<ns1:userPermissions>http://fairspace.ci.fairway.app/iri/user-iri Manage
</ns1:userPermissions>
<ns1:accessMode>Restricted</ns1:accessMode>
<ns1:availableStatuses>Active</ns1:availableStatuses>
<ns1:canDelete>FALSE</ns1:canDelete>
<ns1:iri>https://fairspace.ci.fairway.app/api/webdav/Demonstration</ns1:iri>
<ns1:canWrite>TRUE</ns1:canWrite>
<ns1:ownedByCode>Demo</ns1:ownedByCode>
<ns1:canManage>FALSE</ns1:canManage>
<ns1:canUndelete>FALSE</ns1:canUndelete>
<ns1:workspacePermissions>http://fairspace.ci.fairway.app/iri/workspace-iri
Write
</ns1:workspacePermissions>
<ns1:createdBy>http://fairspace.ci.fairway.app/iri/user-iri</ns1:createdBy>
<ns1:comment>Demonstration collection</ns1:comment>
<ns1:availableAccessModes>Restricted</ns1:availableAccessModes>
<ns1:ownedBy>http://fairspace.ci.fairway.app/iri/workspace-iri</ns1:ownedBy>
<ns1:status>Active</ns1:status>
<d:getcreated>2021-02-02T12:12:33Z</d:getcreated>
<d:creationdate>2021-02-02T12:12:33Z</d:creationdate>
<d:getcontenttype>text/html</d:getcontenttype>
<d:getetag>"https://fairspace.ci.fairway.app/api/webdav/Demonstration"</d:getetag>
<d:iscollection>TRUE</d:iscollection>
<d:displayname>Demonstration collection</d:displayname>
<d:isreadonly>FALSE</d:isreadonly>
<d:name>Demonstration collection</d:name>
<d:supported-report-set></d:supported-report-set>
<d:resourcetype>
<d:collection/>
</d:resourcetype>
</d:prop>
<d:status>HTTP/1.1 200 OK</d:status>
</d:propstat>
</d:response>
</d:multistatus>Create collection or directory
MKCOL /api/webdav/{path} |
|
|---|---|
Create collection or directory |
|
Request headers: |
|
|
Specify the identifier of the owner workspace when creating a collection. |
Example create collection or directory (Python)
import logging
import os
from requests import Request, Response, Session
log = logging.getLogger()
server_url = os.environ.get('FAIRSPACE_URL')
def mkdir(path: str, workspace_iri: str=None):
# Create directory
headers = {
'Authorization': auth()
}
if workspace_iri is not None:
headers['Owner'] = workspace_iri
req = Request('MKCOL', f'{server_url}/api/webdav/{path}/', headers=headers, cookies=self.session().cookies)
response: Response = Session().send(req.prepare())
if not response.ok:
raise Exception(f"Error creating directory '{path}': {response.status_code} {response.reason}")Example create collection or directory (curl)
# Create a new collection, owned by workspace WORKSPACE_IRI
NEW_COLLECTION=New collection
WORKSPACE_IRI=http://fairspace.ci.fairway.app/iri/workspace-iri
curl -i -H "Authorization: ${AUTH_HEADER}" -X MKCOL -H "Owner: ${WORKSPACE_IRI}" "${FAIRSPACE_URL}/api/webdav/${NEW_COLLECTION}"
# Create a new directory in the newly created collection
curl -i -H "Authorization: ${AUTH_HEADER}" -X MKCOL "${FAIRSPACE_URL}/api/webdav/${NEW_COLLECTION}/Test directory"Upload files
|
|
Request data: |
|
|
|
|
Send files with the target file names as keys, see the examples below. |
Example uploading files (Python)
import logging
import os
from requests import Response, Session
log = logging.getLogger()
server_url = os.environ.get('FAIRSPACE_URL')
def upload_files(path: str, files: Dict[str, any]):
# Upload files
response: Response = Session().post(f'{server_url}/api/webdav/{path}/',
headers={'Authorization': auth()},
data={'action': 'upload_files'},
files=files)
if not response.ok:
raise Exception(f"Error uploading files into '{path}': {response.status_code} {response.reason}")Example uploading files (curl)
# Upload files 'coffee.jpg' and 'coffee 2.jpg' to a collection
path="new collection"
curl -i -H "Authorization: ${AUTH_HEADER}" -X POST -F 'action=upload_files' -F 'coffee.jpg=@coffee.jpg' -F 'coffee 2.jpg=@coffee 2.jpg'"${FAIRSPACE_URL}/api/webdav/${path}"Copy and move a directory or file
COPY /api/webdav/{path} |
|
|---|---|
Copy a directory or file. Metadata linked to the file/directory is not copied. |
|
Request headers: |
|
|
The destination path relative to the server, URL encoded, e.g., |
Example copy path (curl)
# Copy 'Examples/Test dir/test 1.txt' to 'Examples/Test dir/test 2.txt'
path="Examples/Test dir/test 1.txt"
target="/api/webdav/Examples/Test%20dir/test%202.txt"
curl -i -H "Authorization: ${AUTH_HEADER}" -X COPY -H "Destination: ${target}" "${FAIRSPACE_URL}/api/webdav/${path}"MOVE /api/webdav/{path} |
|
|---|---|
Move or rename a directory or file. Metadata linked to the file/directory is also moved along with it. |
|
Request headers: |
|
|
The destination path relative to the server, URL encoded, e.g., |
Example move path (curl)
# Move 'Examples/Test dir/test 1.txt' to 'Examples/Test dir/test 2.txt'
path="Examples/Test dir/test 1.txt"
target="/api/webdav/Examples/Test%20dir/test%202.txt"
curl -i -H "Authorization: ${AUTH_HEADER}" -X MOVE -H "Destination: ${target}" "${FAIRSPACE_URL}/api/webdav/${path}"Undelete a directory or file
|
|
Undelete a directory or file |
|
Request headers: |
|
|
|
Request data: |
|
|
|
Example undelete path (curl)
curl -i -H "Authorization: ${AUTH_HEADER}" -X POST -F "action=undelete" "${FAIRSPACE_URL}/api/webdav/${path}"Delete directory content
|
|
Delete directory content |
|
Request data: |
|
|
|
Example delete all in directory (curl)
curl -i -H "Authorization: ${AUTH_HEADER}" -X POST -F "action=delete_all_in_directory" "${FAIRSPACE_URL}/api/webdav/${path}"Revert to a file version
|
|
Restore a previous file version |
|
Request data: |
|
|
|
|
The version number to restore. |
Example revert file version (curl)
curl -i -H "Authorization: ${AUTH_HEADER}" -X POST -F "action=revert" -F "version=${version}" "${FAIRSPACE_URL}/api/webdav/${path}"Other collection actions
On collections, a number of actions is available. These are not documented here in detail, but can be used from the user interface instead.
| Action | Description |
|---|---|
|
Change the access mode of a collection. |
|
Change the status of a collection. |
|
Change the permission of the specified user or workspace on a collection. |
|
Transfer ownership of a collection to another workspace. |
|
Unpublish a published collection. |
SPARQL
The SPARQL API is a standard API for querying RDF databases. This endpoint is read-only and can be used for advanced search, analytics, data extraction, etc. It is only accessible for users with the canQueryMetadata role.
POST /api/rdf/query |
|---|
Execute SPARQL query |
Request body: |
The SPARQL query. |
Example SPARQL query (Python)
Query for the first 500 samples.
import logging
import os
from requests import Response, Session
log = logging.getLogger()
server_url = os.environ.get('FAIRSPACE_URL')
def query_sparql(query: str):
headers = {
'Authorization': auth(),
'Content-Type': 'application/sparql-query',
'Accept': 'application/json'
}
response: Response = Session().post(f"{server_url}/api/rdf/query", data=query, headers=headers)
if not response.ok:
raise Exception(f'Error querying metadata: {response.status_code} {response.reason}')
return response.json()
query_sparql("""
PREFIX example: <https://example.com/ontology#>
PREFIX fs: <https://fairspace.nl/ontology#>
SELECT DISTINCT ?sample
WHERE {
?sample a example:BiologicalSample .
FILTER NOT EXISTS { ?sample fs:dateDeleted ?anyDateDeleted }
}
# ORDER BY ?sample
LIMIT 500
""")Example SPARQL query (curl)
Query for the first 500 samples.
curl -X POST -H "Authorization: ${AUTH_HEADER}" -H 'Content-Type: application/sparql-query' -H 'Accept: application/json' \
-d "
PREFIX example: <https://example.com/ontology#>
PREFIX fs: <https://fairspace.nl/ontology#>
SELECT DISTINCT ?sample
WHERE {
?sample a example:BiologicalSample .
FILTER NOT EXISTS { ?sample fs:dateDeleted ?anyDateDeleted }
}
# ORDER BY ?sample
LIMIT 500
" \
"${FAIRSPACE_URL}/api/rdf/query"RDF metadata
For reading and writing metadata to the database,
the /api/metadata endpoint supports a number of operations:
-
GET: Retrieve metadata for a specified subject, predicate or object. -
PUT: Add metadata -
PATCH: Update metadata -
DELETE: Delete specified triples or all metadata linked to a subject.
The metadata is stored as subject-predicate-object triples. The API supports several serialisation formats for sending :
-
Turtle (
text/turtle) -
JSON-LD (
application/ld+json, JSON schema) -
N-Triples (
application/n-triples)
After any update, the metadata must be consistent with the data model, see Data model and view configuration.
If an update would violate the data model constraints,
the request is rejected with a status 400 response, with a message indicating the violation.
Uploading metadata
Shared metadata entities will in most cases come from other systems and will be added to Fairspace exclusively by an ETL process which will extract data from the laboratory and clinical systems, perform pseudonymization of identifiers, convert the metadata to some RDF-native format conforming the data model and send them to Fairspace.
Fairspace will validate the uploaded metadata against the constraints defined in the data model and returns a detailed error message in case of violations. The validations include all the necessary type checks, referential consistency (validity of identifiers) checks, validation of mandatory fields, etc. If any entity violates the constraints, the entire bulk upload will be rejected.
The ETL process will use a special technical account with the Add shared metadata role. Regular users will not be able to add or modify shared metadata entities. Regular users can link files to shared metadata entities, see Metadata forms and Metadata upload.
In addition to the main ETL workflow, data managers needs a possibility to add or modify certain properties of top-level metadata entities. This can be done using the RDF-based metadata API.
A number of guidelines for uploading shared metadata:
-
Entities must have a type, a globally unique identifier, and a unique label for the type.
It is advised to use a unique identifier from an existing reference system for this purpose. -
Because of the nature of linked data, it is advised to add shared metatdata entities in an append-only fashion: only adding entities and avoid updating or deleting entities.
-
By nature of RDF, metadata is typically added on the level of triples. E.g., when adding a property
dcat:keywordto a file, this will add a key word to the (possibly) already existing list of key words.
If you want to completely replace (or remove) a property from an entity, use thePATCHmethod instead ofPUT.
Example metadata file in turtle format: testdata.ttl:
@prefix example: <https://example.com/ontology#> .
@prefix rdfs: <http://www.w3.org/2000/01/rdf-schema#> .
@prefix subject: <http://example.com/subjects#> .
@prefix file: <http://example.com/api/webdav/> .
@prefix gender: <http://hl7.org/fhir/administrative-gender#> .
@prefix ncbitaxon: <https://bioportal.bioontology.org/ontologies/NCBITAXON/> .
@prefix dcat: <http://www.w3.org/ns/dcat#> .
subject:s1 a example:Subject ;
rdfs:label "Subject 1" ;
example:isOfSpecies ncbitaxon:9606 .
file:coll1\/coffee.jpg
dcat:keyword "fairspace", "java" ;
example:aboutSubject example:s1 .Example uploading metadata file using Python.
import logging
import os
from requests import Response, Session
log = logging.getLogger()
server_url = os.environ.get('FAIRSPACE_URL')
with open('testdata.ttl') as testdata:
response: Response = Session().put(f"{server_url}/api/metadata/",
data=testdata.read(),
headers={
'Authorization': auth(),
'Content-type': 'text/turtle'
})
if not response.ok:
raise Exception(f"Error uploading metadata: {response.status_code} {response.reason}")Example uploading metadata file (curl).
curl -v -X PUT -H "Authorization: Basic $(echo -n "${KEYCLOAK_USERNAME}:${KEYCLOAK_PASSWORD}" | base64)" \
-H "Content-type: text/turtle" --data @testdata.ttl "${FAIRSPACE_URL}/api/metadata/"API specification
GET /api/metadata |
||
|---|---|---|
Retrieve metadata |
||
Parameters: |
||
|
string |
IRI of the subject to filter on. |
|
string |
The predicate to filter on, not required. |
|
string |
The object to filter on, not required. |
|
boolean |
If set, the response will include several properties for the included objects.
The properties to be included are marked with |
Response: |
||
Returns serialised triples matching the query parameters. |
||
Example of fetching metadata in turtle format (curl)
Request metadata for a subject 'a'.
curl -G -H "Accept: text/turtle" \
--data-urlencode "subject=a" \
--data-urlencode "withValueProperties=true" \
"http://localhost:8080/api/metadata/"Example of fetching metadata in json-ld format (curl)
Request metadata for the triple with subject 'a', predicate 'b' and object 'c'.
curl -G -H "Accept: application/ld+json" \
--data-urlencode "subject=a" \
--data-urlencode "predicate=b" \
--data-urlencode "object=c" \
--data-urlencode "withValueProperties=true" \
"http://localhost:8080/api/metadata/"PUT /api/metadata |
||
|---|---|---|
Add metadata. Existing metadata is left untouched.
The data must be consistent with the data model after the update (see Data model and view configuration),
otherwise |
||
Parameters: |
||
|
boolean |
Flag to switch on and off materialized views refresh once metadata updated. The materialized views are used to speed up the metadata search. |
Request body: |
||
Serialised RDF triples. |
||
Example of adding metadata in turtle format (curl)
curl -X PUT -H "Content-type: text/turtle" -d \
'
@prefix example: <https://example.com/ontology#> .
@prefix rdfs: <http://www.w3.org/2000/01/rdf-schema#> .
@prefix test: <https://test.com/ontology#> .
example:Study_001 a test:Study ;
rdfs:label "Project study #001" ;
test:studyIdentifier "STUDY-001" ;
test:studyTitle "Project study #001" ;
test:studyDescription "This is a description of the study." .
' \
"http://localhost:8080/api/metadata/"PATCH /api/metadata/ |
||
|---|---|---|
Update metadata.
Any existing metadata for a given subject/predicate combination will be overwritten with the provided values.
The data must be consistent with the data model after the update (see Data model and view configuration),
otherwise |
||
Parameters: |
||
|
boolean |
Flag to switch on and off materialized views refresh once metadata updated. The materialized views are used to speed up the metadata search. |
Request body: |
||
Serialised RDF triples. |
||
Example of updating metadata in turtle format (curl)
curl -X PATCH -H "Content-type: text/turtle" -d \
'
@prefix example: <https://example.com/ontology#> .
@prefix test: <https://test.com/ontology#> .
example:Study_001 a test:Study ;
test:studyTitle "Updated project study #001" ;
' \
"http://localhost:8080/api/metadata/"DELETE /api/metadata/ |
||
|---|---|---|
Delete metadata. If a request body is provided, the triples specified in the body will be deleted. Otherwise, the subject specified in the subject parameter will be marked as deleted. Please note that the subject will still exist in the database. Only available for users with Add shared metadata role. |
||
Parameters: |
||
|
string |
The subject to filter on. (Optional) |
|
boolean |
Flag to switch on and off materialized views refresh once metadata updated. The materialized views are used to speed up the metadata search. |
Request body: |
||
Serialised RDF triples. (Optional) |
||
Example of deleting triples in turtle format (curl)
curl -X DELETE -H "Content-Type: text/turtle" -d \
'
@prefix example: <https://example.com/ontology#> .
@prefix test: <https://test.com/ontology#> .
example:Study_001 a test:Study ;
test:studyDescription "This is a description of the study." .
' \
"http://localhost:8080/api/metadata/"Example of marking an entity as deleted (curl)
curl -X DELETE -G --data-urlencode "subject=https://example.com/ontology#tpe1" "http://localhost:8080/api/metadata/"Metadata views
Metadata views endpoint used for metadata-based search.
GET /api/views/ |
|---|
List all views with available columns per each view. |
Example list view (curl)
curl -H "Accept: application/json" "http://localhost:8080/api/views/"POST /api/views/ |
||
|---|---|---|
Fetch page of rows of a view matching the request filters. |
||
Parameters: |
||
|
string |
Name of the view. |
|
List of filters, based on available facets and their values. Each filter has to contain a "field" property, matching the name of a facet, and list of values to filter on. |
|
|
integer |
Requested page |
|
integer |
Page size |
Example fetching page of view rows (curl)
curl -X POST -H 'Content-type: application/json' -H 'Accept: application/json' -d \
'{
"view":"Resource",
"filters":[
{
"field":"Resource_type",
"values":["https://fairspace.nl/ontology#Collection"]
}
],
"page":1,
"size":100
}' \
"http://localhost:8080/api/views/"POST /api/views/count |
||
|---|---|---|
Count rows of a view matching request filters. If |
||
Parameters: |
||
|
string |
Name of the view. |
|
List of filters, based on available facets and their values. Each filter has to contain a "field" property, matching the name of a facet, and list of values to filter on. |
|
Example counting view rows (curl)
curl -X POST -H 'Content-type: application/json' -H 'Accept: application/json' -d \
'{
"view":"Resource",
"filters":[
{
"field":"Resource_type",
"values":["https://fairspace.nl/ontology#Collection"]
}
]
}' \
'http://localhost:8080/api/views/count'GET /api/views/facets |
|---|
List all facets with available values per each facet. |
Example retrieving facets with values (curl)
curl -H "Accept: application/json" "http://localhost:8080/api/views/facets"Text search
Search endpoint used for text search on labels or comments.
POST /api/search/files |
||
|---|---|---|
Find files, directories or collections based on a label or a comment. |
||
Parameters: |
||
|
string |
Text fragment to search on. |
|
string |
IRI of the parent directory or collection to limit the search area. |
Response |
||
Object in JSON format, with |
||
|
string |
File (or directory) identifier (IRI). |
|
string |
File (or directory) name. |
|
string |
Type of the resource as defined in the vocabulary, e.g. "https://fairspace.nl/ontology#File", "https://fairspace.nl/ontology#Directory" |
|
string |
File (or directory) description. Optional. |
Example text search (curl)
curl -X POST -H 'Content-type: application/json' -H 'Accept: application/json' -d \
'{
"query":"test folder",
"parentIRI":"http://localhost:8080/api/webdav/dir1"
}' \
'http://localhost:8080/api/search/files'Example text search response
{
"results": [
{
"id": "https://fairspace.example.com/api/webdav/col1/test",
"label": "test",
"type": "https://fairspace.nl/ontology#File",
"comment": "Description of the test file from col1."
},
{
"id": "https://fairspace.example.com/api/webdav/col2/new_test_folder",
"label": "new_test_folder",
"type": "https://fairspace.nl/ontology#Directory",
"comment": null
}
],
"query": "test"
}POST /api/search/lookup |
||
|---|---|---|
Metadata entities lookup search by entity labels or description. |
||
Parameters: |
||
|
string |
Text fragment to search on. |
|
string |
Type of the entity in request. |
Example lookup search (curl)
curl -X POST -H 'Content-type: application/json' -H 'Accept: application/json' -d \
'{
"query":"test",
"resourceType":"https://example.com/ontology#TumorPathologyEvent"
}' \
'http://localhost:8080/api/search/lookup'Workspace management
Operations on workspace entities.
GET /api/workspaces/ |
|
|---|---|
List all available workspaces. |
|
Response contains the following data: |
|
|
Unique workspace IRI. |
|
Unique workspace code. |
|
Workspace title. |
|
List of workspace managers. |
|
Short summary on the workspace - how many collections and how many users it has. |
|
If a current user is added to the workspace as a collaborator. |
|
If a current user is a workspace manager. |
Example of listing available workspaces (curl)
curl -H "http://localhost:8080/api/workspaces/"PUT /api/workspaces/ |
||
|---|---|---|
Add a workspace. Available only to administrators. |
||
Parameters: |
||
|
string |
Unique workspace code. |
Response: |
||
Response contains the workspace name and newly assigned IRI. |
||
Example of adding a workspace (curl)
curl -X PUT -H "Content-type: application/json" -d '{"name": "test workspace"}' "http://localhost:8080/api/workspaces/"DELETE /api/workspaces/ |
||
|---|---|---|
Delete a workspace. Available only to administrators. |
||
Parameters: |
||
|
string |
Workspace IRI (URL-encoded). |
Example of deleting a workspace (curl)
curl -X DELETE --data-urlencode "workspace=http://fairspace.com/iri/123" "http://localhost:8080/api/workspaces/"Workspace users
GET /api/workspaces/users/ |
||
|---|---|---|
List all workspace users with workspace roles. |
||
Parameters: |
||
|
string |
Workspace IRI (URL-encoded). |
Response: |
||
Response contains list of workspace users with their workspace roles. |
||
Example of listing workspace users (curl)
curl -G --data-urlencode "workspace=http://fairspace.com/iri/123" "http://localhost:8080/api/workspaces/users/"PATCH /api/workspaces/users/ |
||
|---|---|---|
Assign a workspace role to a user ( |
||
Parameters: |
||
|
string |
Workspace IRI. |
|
string |
User IRI |
|
string |
|
Example of updating workspace users (curl)
curl -X PATCH -H "Content-type: application/json" -d '{"workspace":"http://fairspace.com/iri/123","user":"http://fairspace.com/iri/456","role":"Member"}' "http://localhost:8080/api/workspaces/users/"Users and permissions
GET /api/users/ |
|---|
List all organisation users. |
Response: |
Returns list of users with user’s unique ID, name, email, username and user’s organisation-level permissions: if a user is an administrator, super-administrator or can view public metadata, view public data or add shared metadata. |
Example listing users (curl)
curl -H 'Accept: application/json' 'http://localhost:8080/api/users/'PATCH /api/users/ |
||
|---|---|---|
Update user roles. |
||
Parameters: |
||
|
string |
UUID of the user for which roles will be updated. |
"role name" |
boolean |
Role name is any of |
Example updating user roles (curl)
curl -X PATCH -H "Accept: application/json" -H "Content-Type: application/json" -d \
'{
"id": "123e4567-e89b-12d3-a456-426614174000",
"canViewPublicData": false,
"canViewPublicMetadata": true
}' \
"http://localhost:8080/api/users/"GET /api/users/current |
|---|
Get current user. |
Response: |
Returns current user’s unique ID, name, email, username and user’s organisation-level permissions: if the user is an administrator, super-administrator or can view public metadata, view public data or add shared metadata. |
Example getting current user (curl)
curl -H "Accept: application/json" "http://localhost:8080/api/users/current"POST /api/users/current/logout |
|---|
logout the current user. |
Example logging out (curl)
curl -X POST "http://localhost:8080/api/users/current/logout"Configuration endpoints
Vocabulary
The vocabulary contains a description of the structure of the metadata. It contains the types of entities that can be created, along with the data types for the fields. It is stored in SHACL format.
GET /api/vocabulary/ |
|---|
Retrieve a representation of the vocabulary. |
Example fetching the vocabulary in turtle format (curl)
curl -H 'Accept: text/turtle' 'http://localhost:8080/api/vocabulary/'Example fetching the vocabulary in json-ld format (curl)
curl -H 'Accept: application/json+ld' 'http://localhost:8080/api/vocabulary/'Features
GET /api/features/ |
|---|
List available application features. |
Response contains list of additional features that are currently available in the application.
Example listing features (curl)
curl -H 'Accept: application/json' 'http://localhost:8080/api/features/'Icons (SVG)
GET /api/iconsvg/{icon_name} |
|---|
Retrieve an SVG icon by name. |
Response contains an SVG icon (if configured).
Example retrieving icon (curl)
curl -H 'Accept: image/svg+xml' 'http://localhost:8080/api/iconsvg/{icon_name}'Services
GET /api/services/ |
|---|
List linked services. |
Response contains list of external services linked to Fairspace,
e.g. JupyterHub, cBioPortal, etc with their configuration details: name, url and icon name that can be used to retrieve
the icon (using GET /api/iconsvg/{icon_name}).
Example listing services (curl)
curl -H 'Accept: application/json' 'http://localhost:8080/api/services/'Server configuration
GET /api/config |
|---|
View server configuration properties. |
Response contains a list of server configuration properties, currently limited to a max file size for uploads.
Example listing properties (curl)
curl -H 'Accept: application/json' 'http://localhost:8080/api/config/'External storages
GET /api/storages/ |
|---|
List linked data storages. |
Response contains list of external data storages linked to Fairspace.
Example listing external storages using curl
curl -H 'Accept: application/json' 'http://localhost:8080/api/storages/'Maintenance
POST /api/maintenance/reindex |
|
|---|---|
Recreate the view database from the RDF database. Starts an asynchronous task to clean the PostgreSQL database with the data used for the metadata views, and to repopulate the database with the data from the RDF database. This can be used after a change in the data model or view configuration to ensure that all data is properly indexed. Only available when the application is configured with Only allowed for administrators. |
|
Response: |
|
|
Asynchronous task to recreate the index has started. |
|
Operation not allowed. The current user is not an administrator. |
|
Maintenance (reindexing or compacting) is already in progress. |
|
Service not available. This means that the application is configured not to use a view database. |
POST /api/maintenance/compact |
|
|---|---|
Compact the Jena TDB database files. Jena database files grow fast when using transactions. This operation will compact the database files to reduce their size. If data is inserted using many small transactions the files will be reduced to 10-20% of their original size. Only allowed for administrators. |
|
Response: |
|
|
Asynchronous task to compact Jena files has started. |
|
Operation not allowed. The current user is not an administrator. |
|
Maintenance (reindexing or compacting) is already in progress. |
|
Service not available. This means that the application is configured not to use a view database. |
Example of compacting Jena using curl
curl -X POST 'http://localhost:8080/api/maintenance/compact'GET /api/maintenance/status |
|
|---|---|
Get the status of maintenance tasks. It is not possible to run more than one maintenance task at the same time. If you start a task while another task is running, the new task will be rejected. If you want to know in advance whether a task is running, you can use this endpoint. A text is return |
|
Response: |
|
|
Returns "active" or "inactive" |
|
Operation not allowed. The current user is not an administrator. |
|
Reindexing is already in progress. |
|
Service not available. This means that the application is configured not to use a view database. |
Example of getting the maintenance status using curl
curl -X POST 'http://localhost:8080/api/maintenance/status'External file system integration
As Fairspace supports the WebDAV protocol, it can be configured to connect to external data storages that implement a WebDAV interface. An overview of external files is integrated into Fairspace user interface. Currently, a read-only interaction is supported. Users can browse through the external file system, read the data and metadata (e.g. creation date, description). Files from the external storage will be also made available for analysis in Jupyter Hub.
Access policy
Access policies differ between systems. To avoid inconsistencies, permissions validation and management are expected to be under control of the external storage system. Each storage component is responsible for its own policy and needs to perform the required checks to ensure that users only get to see the data they are supposed to see.
It is assumed that a user requesting files from a storage using WebDAV has at least "read" access to all the files included in the WebDAV response.
Access can be further limited by using a custom access property. If a value of this property on a resource is set to "List",
the resource’s metadata will be readable, but it will not be possible to read the resource’s content.
Another assumption is that the Fairspace client can authenticate in the external storage via the same Keycloak and the same realm as configured for Fairspace, so that the same bearer token can be used for all storages. See the Authentication section for more information.
API specification and supported parameters
A subset of default WebDAV properties is used and displayed as a resource metadata in the Fairspace user interface. These properties are presented in the table below.
| WebDAV property | Description |
|---|---|
|
Creation date |
|
Flag determining whether a resource is a file or directory |
|
Last modification date |
|
Size of the file (0 for directories) |
There is also a set of custom Fairspace properties, some of which are required to be returned from the WebDAV request.
| WebDAV property | Description |
|---|---|
|
IRI of the resource. Required. |
|
Id of a user that created the resource. |
|
Resource description. |
|
By default, users are granted |
|
List of IRIs in a form of comma-separated string. IRIs represent all metadata entities linked to the resource. If the IRI matches a metadata entity stored in Fairspace, such an entity will be displayed in the user interface. |
It is also supported to specify any other custom property in the WebDAV response body, as WebDAV responses are easily extendable. All these properties (if not specifically marked as excluded in Fairspace), will be displayed in the user interface in a form of key-value pairs.
Text search in external storages
Text based search on external file system can be enabled in the Fairpsace user interface,
if the external system exposes a search endpoint, following the specification from the Text search section.
To enable finding files based on name or description, searchUrl has to be specified in the storage configuration.
Configuration
Multiple external storages can be configured simultaneously. A list of configuration parameters is presented below.
| Parameter | Description |
|---|---|
|
Unique name of the storage. |
|
String to be used as a display name of the storage. |
|
WebDAV endpoint to connect to. |
|
Optional search endpoint URL. If specified, a text based search on file name or description will be enabled in the user interface. |
|
Optional IRI of the root directory. If not specified, |
Sample configuration of storages in YAML format:
storages:
exStorage1:
name: exStorage1
label: "External storage 1"
url: https://exstorage1/api/webdav
searchUrl: https://exstorage1/api/search/
rootDirectoryIri: http://ex1/api/webdav/
exStorage2:
name: exStorage2
label: "External storage 2"
url: https://exstorage2/api/webdavMulti-Fairspace metadata views integration
Fairspace works with a single data model configured. However, if there is a need to have data from multiple models, representing multiple domains that would be combined in single user interface, it is possible to integrate multiple Fairspace instances together.
The main Fairspace instance can be configured as a view interface of metadata from another (external) Fairspace instance. External Fairspace can have a different metadata model configured than the main Fairspace. An important precondition is that the external instance has to be connected to the same Keycloak realm as the main instance.
External metadata can be searched and browsed in the same way as the internal metadata views. When configured, there is an extra page, separate from internal metadata views, that allows to explore external metadata. Currently, cross-instance metadata search is not supported.
For the external metadata pages, the tables and columns are created basing on the views configuration specified in that instance configuration.
Access policy
Connection to the same Keycloak realm allows to authenticate a user in all integrated Fairspace instances with a single set of login credentials (single sign-on). Each Fairspace instance is responsible for controlling the access to its own metadata and perform the required checks to ensure that users only get to see metadata of an instance, if has a view public metadata role assigned within that instance.
Configuration
Multiple external Fairspace metadata pages can be configured simultaneously. A list of configuration parameters is presented below.
| Parameter | Description |
|---|---|
|
Unique name of the metadata source. |
|
String to be used as a display name of the metadata source. |
|
Fairspace instance to connect to. If the url is not specified, the metadata source will be treated as the internal one. Important! There should only be a single configuration of internal metadata (only the first one will not be ignored). |
|
Name of an icon configured in the "icons" section of values.yaml file. If the name is not specified, there will be a default icon used. |
Sample configuration of external metadata sources in YAML format:
fairspace:
...
metadata-sources:
internal:
name: internal
label: "Internal metadata"
icon-name: "icon-internal-metadata"
additionalMetaSource1:
name: metadataSource1
label: "Test metadata 1"
url: https://fairspace-test1/api/
icon-name: "icon-1"
metaSource2:
name: metadataSource2
label: "Test metadata 2"
url: https://fairspace-test2/api/Configuration of gateway redirection in values.yaml:
pluto:
...
backends:
storageRoutes:
- id: additional-metadata-domain1
uri: https://fairspace-test1
predicates:
- Path=/api/metadata-sources/metadataSource1/**
filters:
- RewritePath=/api/metadata-sources/metadataSource1/(?<segment>(views|vocabulary|metadata).*),/api/$\{segment}
- RewriteResponseHeader=Set-Cookie, ^([^=]+)=, DO_$1=
- id: additional-metadata-domain2
uri: https://fairspace-test2
predicates:
- Path=/api/metadata-sources/metadataSource2/**
filters:
- RewritePath=/api/metadata-sources/metadataSource2/(?<segment>(views|vocabulary|metadata).*),/api/$\{segment}
- RewriteResponseHeader=Set-Cookie, ^([^=]+)=, DO_$1=where RewriteResponseHeader is important filter that needs to be added not to overwrite an existing browser session, which would lead to authorization errors.
Configuration above gets transformed to the following Spring Cloud Gateway config in the application.yaml configuration of Pluto:
spring:
cloud:
gateway:
routes:
- id: additional-metadata-domain1
uri: https://fairspace-test1
predicates:
- Path=/api/metadata-sources/metadataSource1/**
filters:
- RewritePath=/api/metadata-sources/metadataSource1/(?<segment>(views|vocabulary|metadata).*),/api/$\{segment}
- RewriteResponseHeader=Set-Cookie, ^([^=]+)=, DO_$1=
- ...Structure and terminology
In this section we describe in detail the main concepts and components of the Fairspace data repository and how they relate to each other.
The core entities of the data repository are:
-
Users: individual users in the organisation, looking for data, contributing to data collections or managing data.
-
Workspaces (for projects, teams): entities in the system linked, representing a group of users, to organise data collections and data access.
-
Collections: entities in the system to group data files. These are the minimal units of data for data access and data modification rules.
-
Files: The smallest units of data that the system processes. Files always belong to a single collection. Files can be added, changed and deleted, but not in all collection states. Changing a file creates a new version. Access to a file is based on access to the collection the file belongs to. Files can be organised in Directories, which we will leave out of most descriptions for brevity.
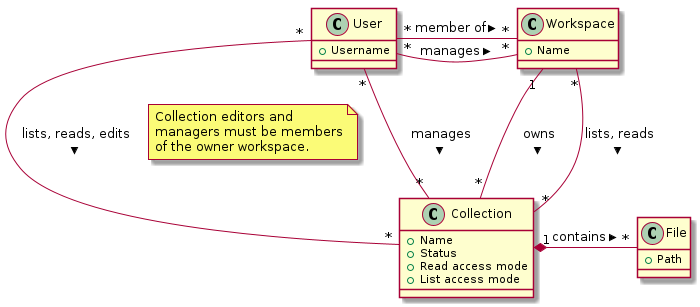
The diagram above sketches the relevant entities and actors. The basic structure consists of users, workspaces, collections and files as represented in the system. Collections are the basic units of data access management. A collection is owned by a workspace. The responsibility for a collection is organised via the owner workspace: members of the owner workspace can be assigned as editors or managers of the collection. This reflects the situation where in an organisation, a data collection belongs to a project or a research team. This way the workspace represents the organisational unit that is responsible for a number of data collections (e.g., a research team or project). Data can be shared with other workspaces or individual users (for reading) and ownership may be transferred to another workspace (e.g., in the case the workspace is temporary, or when the organisation changes).
Fairspace provides a data catalogue, containing all the metadata, which is visible for all users with catalogue access (View public metadata). Users with metadata write access (Add shared metadata) can add metadata to the catalogue. Preferably this is done by an automated process that ensures the consistency of the metadata and uniqueness of metadata entities. Metadata on collection and file level is protected by the access policy of the collections.
User administration is organised in an external component ([Keycloak]), but user permissions are stored in Fairspace. A back end application is responsible for storing the data and metadata, and for providing APIs for securely retrieving and adding data and metadata using standard data formats and protocols. A user interface application provides an interactive file manager and (meta)data browser and data entry forms based on the back end APIs. Besides the data storage and data management, Fairspace offers analysis environments using Jupyter Hub. In Jupyter Hub, the data repository is accessible. Every user has a private working directory. We do no assumptions on the structure of the data or on the permissions of the external file systems that are connected to the data repository and referenced in the data catalogue. The organisation structure may be replicated in the different systems in incompatible ways, and the permissions may not be aligned.
Workflow and access modes
During the lifetime of a collection, different rules may be applicable for data modification and data access. In Fairspace, collections follow a workflow with the following statuses:
-
Active: for the phase of data collection, data production and data processing;
-
Read-only: for when the data set is complete and is available for reuse;
-
Archived: for when the data set should not be available for reading, but still needs to be preserved;
-
Deleted: for when the data set needs to be permanently made unavailable (non-readable and non-searchable). This status is irreversible. There is one exception to this rule – for the sake of data loss prevention, in special cases, administrators can still undelete a collection that was already deleted.
In these different statuses, different actions on the data are enabled or disabled. Also, visibility of the data and linked metadata depends partly on the collection status. We also distinguish three access modes for reading and listing files in a collection (where listing also includes seeing the metadata):
-
Restricted: only access to explicitly selected workspaces and users;
-
Metadata published: the collection and its files are visible, metadata linked to them is visible for all users;
-
Data published: the files in the collection are readable for all users. This mode is irreversible. There is one exception to this rule – there might be a special situation, resulting from, e.g., a legal reason, when a collection has to be unpublished. This action is available to administrators, but it is highly discouraged, since the collection (meta)data may already be referenced in other systems.
The statuses and access modes, and the transitions between them are shown in the following diagram.
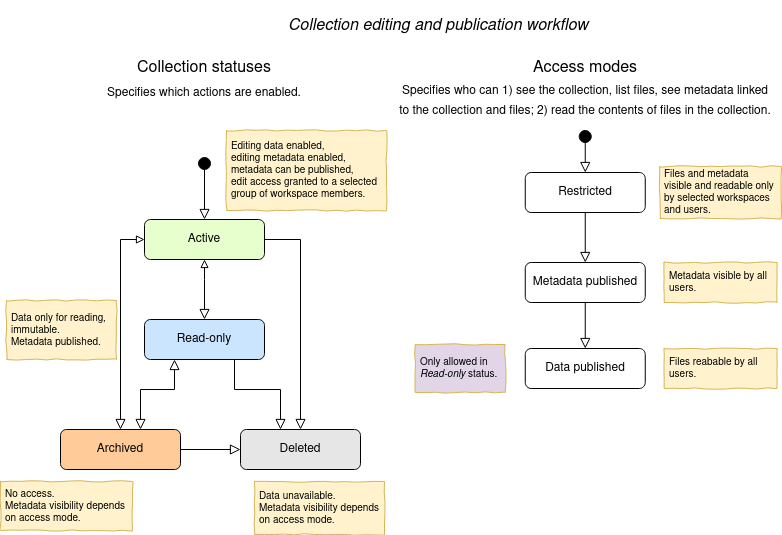
Roles and permissions
We distinguish the following roles in the solution:
-
User: regular users can only view their own workspaces and collections.
-
View public metadata: the user can view public metadata, workspaces, collections and files;
-
View public data: the user can read public files;
-
Admin: can create workspaces, assign roles and permissions;
-
Add shared metadata: can add, modify and delete shared metadata entities.
-
Query metadata: can run SPARQL queries to query metadata.
Most users should have the View public data role. Only when the shared metadata may contain sensitive information that should not be visible for some users, the public data and public metadata roles should be discarded for those users.
Workspaces are used to organise collections in a hierarchy. On workspace level there are two access levels:
-
Manager: can edit workspace details, manage workspace access and manage access to all collections that belong to the workspace;
-
Member: can create a collection in the workspace.
Access to collections and files is managed on collection level. We distinguish the following access levels on collections:
-
List: see collection, directory and file names and metadata properties/relations (only applicable for collections shared via the Metadata published access mode);
-
Read: read file contents;
-
Write: add files, add new file versions, mark files as deleted;
-
Manage: grant, revoke access to the collection, change collection status and modes.
Access levels are hierarchical: the Read level includes the List level; the Edit level includes Read level; the Manage level includes Edit and Read level access. The user that creates the collection gets Manage access.
Data model and view configuration
Metadata
Metadata is data about data. Metadata is used to describe data assets, e.g., for making it easier to find or use certain data. Because metadata is data itself, it can be difficult to make a proper distinction between data and metadata in a system.
Types of metadata
In a digital archive, technical metadata is linked to data assets, like file type, location, size, creation or modification dates, checksums for checking data integrity, ownership.
Such metadata is essential for a system to store and retrieve data files.
Technical metadata can also include data format specific properties, like encoding, data layout, resolution, etc., required to correctly read the data.
With most publications, bibliographic metadata is associated, such as author, title, abstract, publication details, keywords and subject categories.
Such metadata makes it possible to find relevant publications.
This is the kind of metadata used by libraries and archives and numerous standards exist for such data, such as Dublin Core and METS.
More detailed descriptive metadata provides information about the contents of the data, e.g., description of rows and columns, summary statistics, project information, geographical information, results, study design, methods, materials or equipment. In the extreme case, the entire content of the file is captured in descriptive metadata.
We can distinguish different kinds of descriptive metadata, such as:
-
Description of the contents (rows, columns, values, summary statistics)
-
Description of the subject, what the data is about (subject, topic, project, study design, object of study, time, location)
-
Description of data sources (for derived or processed data)
-
Description of the methods or technology used to produce or capture the data, such as scripts and versions.
In the context of health research data, it is essential to link data to research subjects, i.e., patients and samples.
The values of the metadata can be of any type, numerical, free text, date, conform to a controlled vocabulary (e.g., ICD or SNOMED codes, units, file types) or a reference to a typed entity within the database, or external entities.
Likewise, the data the metadata is about can be of any type, a file system, a tabular file, image, genomic data, a relational database, etc.
Purpose
Metadata is used for several purposes:
-
Descriptors to enable use of the data (file type, file format, encoding, how it was created/generated). The metadata may be used by users or scripts to read or interpret a particular file or data set.
-
Finding relevant data for analysis:
-
Metadata may be used to organise data within a data set that a researcher is working on, by using (study specific) categories linked to individual files.
-
Metadata may be used in search queries or navigation to find out if data is available that meets certain selection criteria (e.g., data types, categories, cohort characteristics), for inclusion in a new analysis.
-
Metadata may be used to identify data that is linked to a specific entity, such as a patient or a sample, to determine if such data has already been analysed, in order to avoid duplicate analysis.
-
It is important to identify for which purpose metadata is collected and used, as it may affect which types of metadata are collected, how they are navigated and if access control on metadata is desired or required.
Data model
To enable validation of (meta)data, and to enable intuitive navigation and search within the metadata, it is essential to have a good data model.
The data model consists of the entity types (classes), their properties (with types) and relationships between entities that can be represented in the system.
The data model needs to be broad (expressive) enough to allow users to express all relevant facts about data sets conveniently and accurately, but it needs to be specific enough to allow validation and the generation of useful overviews and information pages. International data standards should be used as much as possible to enable interoperability between systems.
E.g., it is probably better to use a specific field ‘disease’ where the value must be a valid ICD-10 code, than using a generic ‘description’ field where a disease is described in a free text field.
Data domains
We distinguish different data domains in order to clearly separate the data that is system specific and the metadata that is more flexible.
Workspaces and collection-level data
Users, workspaces, collections, directories and files are system-level entities, representing the file system of the system. Access to these entities is restricted by the workspace-level and collection-level access control. These entities cannot be changed on demand, but are inherent to the system. However, custom properties and relations may be added, e.g., to link files to patients.
Metadata
The data model for the other (non system-level) entities, the shared metadata, can be configured, in order to make the metadata suitable for the environment where it is used. These metadata are used to link entities in the file system to entities in the research domain, such as samples, patients, diseases, diagnoses, or to entities in the organisation domain, such as projects. These entities may be displayed and navigated in the application and can be explored through the API (for technical users).
Controlled vocabularies
The data model may contain controlled vocabularies (e.g., disease codes, file types, project phases) that can be used as values in the metadata. Every value in a controlled vocabulary has a unique identifier and a label. Using such vocabularies enables standardisation and validation of metadata values.
Reference data
The data model may support domain specific entity types (patients, samples, genes, treatments, studies, etc.) or generic entity types (project, organisation, person, etc), defining the metadata objects that collection-level data assets can refer to. The reference data can also be linked.
Every entity has a unique identifier, a type, a label, and the properties and relations as specified by the type. These entities do not belong to a particular space that is owned by a specific group or user.
Data model configuration
Fairspace uses an Apache Jena database to store system metadata and the custom domain specific metadata. The data models for these metadata are defined using the Shapes Constraint Language (SHACL).
-
The system metadata includes workspaces, collections, directories, files, file versions, users and access rights. The system data model is defined in system-vocabulary.ttl
-
The customisable data model includes the custom (shared) metadata entities, custom controlled vocabulary types, and custom properties of the system entities. The default custom data model is defined in vocabulary.ttl. This data model can be overriden by a data more suitable for your organisation.
A schematic overview of the default data model in vocabulary.ttl:
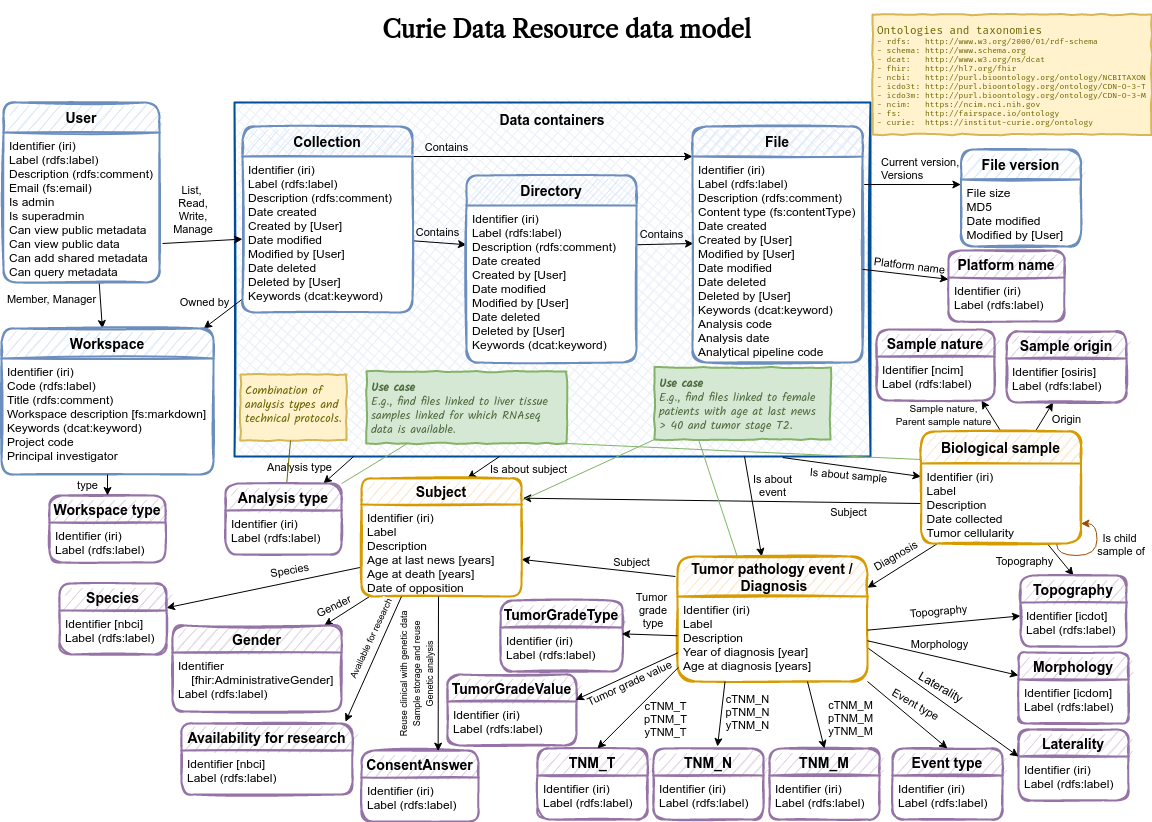
The data model defines an entity-relationship model, specifying the entity types that are relevant to describe your data assets, the properties of the entities, and the relationships between entities.
In this example data model, the following custom entity types are defined:
-
example:Genderwith property Label; -
example:Specieswith property Label; -
example:Subjectwith properties Gender, Species, Age at last news and Files.
The system class fs:File is extended with the Is about subject property.
@prefix owl: <http://www.w3.org/2002/07/owl#> .
@prefix rdf: <http://www.w3.org/1999/02/22-rdf-syntax-ns#> .
@prefix rdfs: <http://www.w3.org/2000/01/rdf-schema#> .
@prefix sh: <http://www.w3.org/ns/shacl#> .
@prefix xsd: <http://www.w3.org/2001/XMLSchema#> .
@prefix dash: <http://datashapes.org/dash#> .
@prefix fs: <https://fairspace.nl/ontology#> .
@prefix example: <https://example.com/ontology#> .
example:Gender a rdfs:Class, sh:NodeShape ;
sh:closed false ;
sh:description "The gender of the subject." ;
sh:name "Gender" ;
sh:ignoredProperties ( rdf:type owl:sameAs ) ;
sh:property
[
sh:name "Label" ;
sh:description "Unique gender label." ;
sh:datatype xsd:string ;
sh:maxCount 1 ;
dash:singleLine true ;
fs:importantProperty true ;
sh:path rdfs:label
] .
example:Species a rdfs:Class, sh:NodeShape ;
sh:closed false ;
sh:description "The species of the subject." ;
sh:name "Species" ;
sh:ignoredProperties ( rdf:type owl:sameAs ) ;
sh:property
[
sh:name "Label" ;
sh:description "Unique species label." ;
sh:datatype xsd:string ;
sh:maxCount 1 ;
dash:singleLine true ;
fs:importantProperty true ;
sh:path rdfs:label
] .
example:isOfGender a rdf:Property .
example:isOfSpecies a rdf:Property .
example:ageAtLastNews a rdf:Property .
example:Subject a rdfs:Class, sh:NodeShape ;
sh:closed false ;
sh:description "A subject of research." ;
sh:name "Subject" ;
sh:ignoredProperties ( rdf:type owl:sameAs ) ;
sh:property
[
sh:name "Label" ;
sh:description "Unique subject label." ;
sh:datatype xsd:string ;
sh:maxCount 1 ;
dash:singleLine true ;
fs:importantProperty true ;
sh:path rdfs:label;
sh:order 0
],
[
sh:name "Gender" ;
sh:description "The gender of the subject." ;
sh:maxCount 1 ;
sh:class example:Gender ;
sh:path example:isOfGender
],
[
sh:name "Species" ;
sh:description "The species of the subject." ;
sh:maxCount 1 ;
sh:class example:Species ;
sh:path example:isOfSpecies
],
[
sh:name "Age at last news" ;
sh:description "The age at last news." ;
sh:datatype xsd:integer ;
sh:maxCount 1 ;
sh:path example:ageAtLastNews
],
[
sh:name "Files" ;
sh:description "Linked files" ;
sh:path [sh:inversePath example:aboutSubject];
] .
example:aboutSubject a rdf:Property .
# Augmented system class shapes
fs:File sh:property
[
sh:name "Is about subject" ;
sh:description "Subjects that are featured in this collection." ;
sh:class example:Subject ;
sh:path example:aboutSubject
] .All entity types have a unique label, specified using the rdfs:label predicate.
The Gender and Species properties link the subject to an entity from
the respective controlled vocabularies.
The Age at last news property is a numerical (integer) value property.
The Files property of the Subject entity type is an example of an inverse relation.
The link is defined on the file, but the link will be visible on the subject as well, because of this inverse relation.
The following guidelines should be followed when creating a custom data model.
-
Define a namespace for your custom entities and properties, like
@prefix example: https://example.com/ontology# .in the example. -
Each custom entity type must have types
rdfs:Classandsh:NodeShape, the propertiessh:closed falseandsh:ignoredProperties ( rdf:type owl:sameAs ), and a valid value forsh:name. Thesh:descriptionproperty is optional. -
Controlled vocabulary or terminology types are modelled as entity types as well, having only the Label (
rdfs:label) property, seeexample:Genderandexample:Species. -
Properties are specified using the
sh:propertyproperty.-
Every entity type must have a property Label (
sh:path rdfs:label) of data typexsd:string. The label of an entity must be unique for that type. The label property should be singleton and markedfs:importantProperty true. If there are multiple properties, the label should havesh:order: 0. -
Properties must have a valid value for
sh:name. Thesh:descriptionproperty is optional. -
A property must either have a
sh:datatypeproperty, specifying one ofxsd:string,xsd:integerorxsd:date, or a propertysh:classspecifying an entity type as the target of a relationship. -
The predicate used for the property (the middle part of the RDF triple) is specified with the
sh:pathproperty, e.g.,example:aboutSubjectfor the Is about subject relation. -
If a relationship is bidirectional, the path of the inverse relation is specified using
sh:inversePath, see the Files property on the Subject entity type. -
A property can be marked mandatory by specifying
sh:minCount 1. A property can be marked singleton by specifyingsh:maxCount 1. -
A text property (with
sh:datatype xsd:string) can be limited to a single line text field usingdash:singleLine true.
-
Limitations
Although assigning multiple types to an entity is easy in RDF, Fairspace assumes entities to have a single type.
Inheritance is possible in SHACL, but not supported by Fairspace. Instead of specifying an entity type as a subtype of another, a single type can be specified with a type property, indicating the sub type of the entity.
E.g., instead of defining entity types DNASeqAssay and RNASeqAssay as sub types of Assay, a property type assayType can be defined on Assay, using a controlled vocabulary type AssayType with the assay types as values.
Although there are many RDF-compatible XSD datatypes, it is recommended to reuse the types
that are already used in the default vocabularies.ttl file as a value of sh:datatype property.
Other types may not be handled properly in the user interface and may cause some unexpected issues.
Same recommendation is for SHACL constraints that can be added for an entity or its properties - reuse the constraints described
in the custom data model creation guidelines.
Changing existing data model
Flexible, configurable data model is one of the key features of Fairspace. Data model evolution is possible, but needs to be applied carefully as well: make sure that new versions of data models are consistent with previous versions, in order to prevent inconsistencies for existing data.
| Editing a data model is specialized work for data modellers/information architects. Use with care. The system is flexible, but the system cannot compensate for poor data modelling choices. Bad modelling will make it hard for users to enter data and to interact with the data. |
It is recommended to only add properties to existing entities or add new entities. Changing existing entities will cause inconsistencies.
List of data model changes that can be considered safe:
-
Adding new entity,
-
Adding new property to an existing entity,
-
Removing constraints on properties,
-
Changing description of an existing entity or property.
Dangerous actions (not recommended):
-
Changing or removing existing entities,
-
Adding or changing constraints,
-
Removing or changing existing properties (property type, name),
-
Changing relations between entities.
-
Update the vocabularies.ttl file, defining the custom model. Follow the guides specified in Data model configuration section).
-
Update views configuration file (see views View configuration section), if applicable - only if there is a change that needs to be reflected in metadata search views.
-
Apply the changes
For the deployment with Helm, run an upgrade command with saturn.vocabulary and saturn.views parameters pointing to a new vocabularies and views definitions (see Installation and configuration), use
--set-fileoption:~bin/helm/helm upgrade … --set-file saturn.vocabulary=/path/to/vocabulary.ttl --set-file saturn.views=/path/to/views.yamlThis should also restart the Saturn pod. If not, trigger the restart manually.
For local development - replace vocabulary file in projects/saturn/vocabulary.ttl and views configuration in projects/saturn/views.ttl. Restart Saturn run.
-
Load data for new entities or properties.
-
Reindex Postgres database using
/api/maintenance/reindexAPI endpoint (see Maintenance API) to apply the changes for metadata search.
Controlled vocabularies
For controlled vocabulary types, e.g., Gender and Species in the example, you should insert the allowed values in the database by uploading a taxonomies file using the RDF metadata API. An example taxonomy is in taxonomies.ttl.
It is preferred to use existing standard taxonomies and labels. If that is not possible, please define your own namespaces for your custom taxonomies.
In this example we use existing standard ontologies for the Gender and Species controlled vocabulary types.
-
The HL7 FHIR AdministrativeGender code system for Gender.
-
The NCBI Organismal Classification for Species.
@prefix rdfs: <http://www.w3.org/2000/01/rdf-schema#> .
@prefix example: <https://example.com/ontology#> .
@prefix gender: <http://hl7.org/fhir/administrative-gender#> .
@prefix ncbitaxon: <https://bioportal.bioontology.org/ontologies/NCBITAXON/> .
gender:male a example:Gender ;
rdfs:label "Male" .
gender:female a example:Gender ;
rdfs:label "Female" .
ncbitaxon:562 a example:Species ;
rdfs:label "Escherichia coli" .
ncbitaxon:1423 a example:Species ;
rdfs:label "Bacillus subtilis" .
ncbitaxon:4896 a example:Species ;
rdfs:label "Schizosaccharomyces pombe" .
ncbitaxon:4932 a example:Species ;
rdfs:label "Saccharomyces cerevisiae" .
ncbitaxon:6239 a example:Species ;
rdfs:label "Caenorhabditis elegans" .
ncbitaxon:7227 a example:Species ;
rdfs:label "Drosophila melanogaster" .
ncbitaxon:7955 a example:Species ;
rdfs:label "Zebrafish" .
ncbitaxon:8355 a example:Species ;
rdfs:label "Xenopus laevis" .
ncbitaxon:9606 a example:Species ;
rdfs:label "Homo sapiens" .
ncbitaxon:10090 a example:Species ;
rdfs:label "Mus musculus" .View configuration
For the metadata pages in the user interface, a view configuration needs to be created that specifies the tables and columns. An example can be found in views.yaml
Installation and configuration
Local development
Requires:
-
yarn
-
docker
-
Java 21
On MacOS the docker logging driver needs to be configured, because the default is not available (journald).
Override the logging driver by setting the DOCKER_LOGGING_DRIVER environment variable or adding a line the .env file in local-development:
DOCKER_LOGGING_DRIVER=json-fileTo run the development version, checkout this repository,
navigate to projects/mercury and run the following commands (yarn install only has to be ran the first time running fairspace).
yarn install
yarn devThis will start a Keycloak instance for authentication at port 5100,
the backend application named Saturn at port 8080 and the
user interface at port 3000.
At first run, you need to configure the service account in Keycloak. If you cannot log in, you might need to restart fairspace by closing it and running yarn dev again.
-
Navigate to http://localhost:5100
-
Login with credentials
keycloak,keycloak -
In the top-left drop down menu, select the fairspace realm
-
Grant the
view-usersrole to the client service account:-
Click
Clientsin the left menu → Select 'workspace-client' -
Choose tab
Service Account Roles -
Click
Assign Role -
Select
Filter by clientsfrom the drop down menu and search for role nameview-users. Then clickAssign.
-
Now everything should be ready to start using Fairspace:
-
Navigate to http://localhost:3000/dev to open the application.
-
Login with one of the following credentials:
Username Password organisation-admin
fairspace123
user
fairspace123
Kubernetes and helm
Requires:
-
Helm >= 3.14.x
-
kubectl >= 1.27.x
You can deploy Fairspace on a Kubernetes cluster using Helm. Helm charts for Fairspace are published to the public helm repository at ghcr.io/thehyve/helm-charts/fairspace (GitHub Packages of the Fairspace repository)
We provide a number of charts for various components that can be used in combination, or separately:
-
Fairspace Keycloak: Installs Keycloak and configures an ingress node for Keycloak. This chart is not required if Keycloak is already installed separately. You still need to configure a Keycloak realm for Fairspace (chart source: https://github.com/thehyve/fairspace-keycloak).
-
Fairspace: Installs the Fairspace application, including the saturn backend, pluto proxy, mercury frontend and a PostgreSQL database, and configures an ingress node for Fairspace (chart source: https://github.com/thehyve/fairspace).
-
Jupyter: Installs a version of Jupyter Hub that uses Keycloak for authentication and launches a jupyter-datascience-notebook based Jupyter notebook with the Fairspace collections file system mounted automatically (chart source: https://github.com/thehyve/fairspace-jupyter).
Instructions for deploying to Google Cloud
Download and install helm and gcloud
-
Download
helm 3.14.3(or higher) from https://github.com/helm/helm/releases/tag/v3.14.3 -
Extract the downloaded archive to
~/bin/helmand check with:~/bin/helm/helm version -
Install kubectl (for Helm 3.14.x install version > 1.27.x).
-
Download and install the Google Cloud SDK (requires Python).
-
Obtain credentials for Kubernetes:
gcloud iam service-accounts keys create credentials.json --iam-account <iam account id, e.g. fairspace-207108@appspot.gserviceaccount.com> export GOOGLE_APPLICATION_CREDENTIALS=/path/to/credentials.json gcloud container clusters get-credentials <cluster id, e.g. fairspacecicluster> --zone europe-west1-b --project <project id, e.g. fairspace-207108> -
Check if all tools are correctly installed:
# List available clusters gcloud container clusters list # List Kubernetes namespaces kubectl get ns # List helm releases (deployments) ~/bin/helm/helm list -A
Configure DNS
Find the address of the Kubernetes cluster:
kubectl cluster-infoCreate DNS records for the keycloak.example.com, fairspace.example.com and (optionally) jupyterhub.example.com domains, pointing to the cluster.
Fetch charts
# Fetch the fairspace-keycloak chart
~/bin/helm/helm pull oci://ghcr.io/thehyve/fairspace/helm-charts/fairspace-keycloak --version 0.7.0
# Fetch the fairspace chart
~/bin/helm/helm pull oci://ghcr.io/thehyve/fairspace/helm-charts/fairspace --version 2.0.2Deploy Keycloak
Create a new Kubernetes namespace:
kubectl create namespace keycloak-newCreate a new deployment (called release in helm terminology) and install the Fairspace Keycloak chart:
~/bin/helm/helm install keycloak-new fairspace-keycloak-0.7.0.tgz --namespace=keycloak-new \
-f /path/to/fairspace-keycloak-values.yamlYou can pass values files with -f or --values.
Example fairspace-keycloak-values.yaml file:
fairspaceKeycloak:
name: keycloak-new
postgresql:
postgresPassword: # choose a strong database password
keycloak:
extraEnv: |
- name: KEYCLOAK_USER
value: keycloak
- name: KEYCLOAK_PASSWORD
value: # choose a strong Keycloak admin password
- name: PROXY_ADDRESS_FORWARDING
value: "true"
ingress:
domain: keycloak.example.com
tls:
certificate:
force: trueYou can pass values files with -f or --values.
Configure a Keycloak realm for Fairspace
-
Navigate to
https://keycloak.example.comand select Administration Console:
-
Create a realm, e.g., fairspace:
-
Configure the realm:
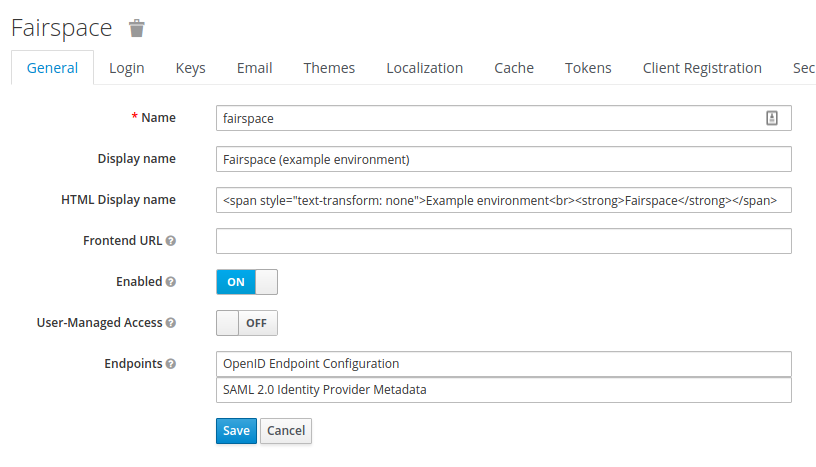
-
Add a client to the realm, e.g., fairspace-example-private:
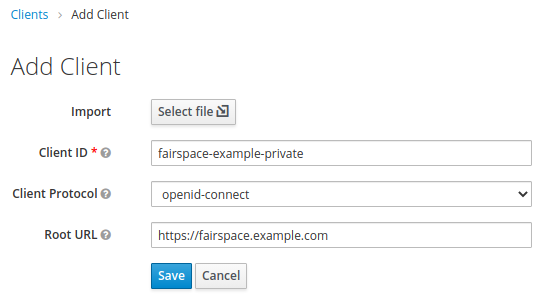
-
Configure the client:
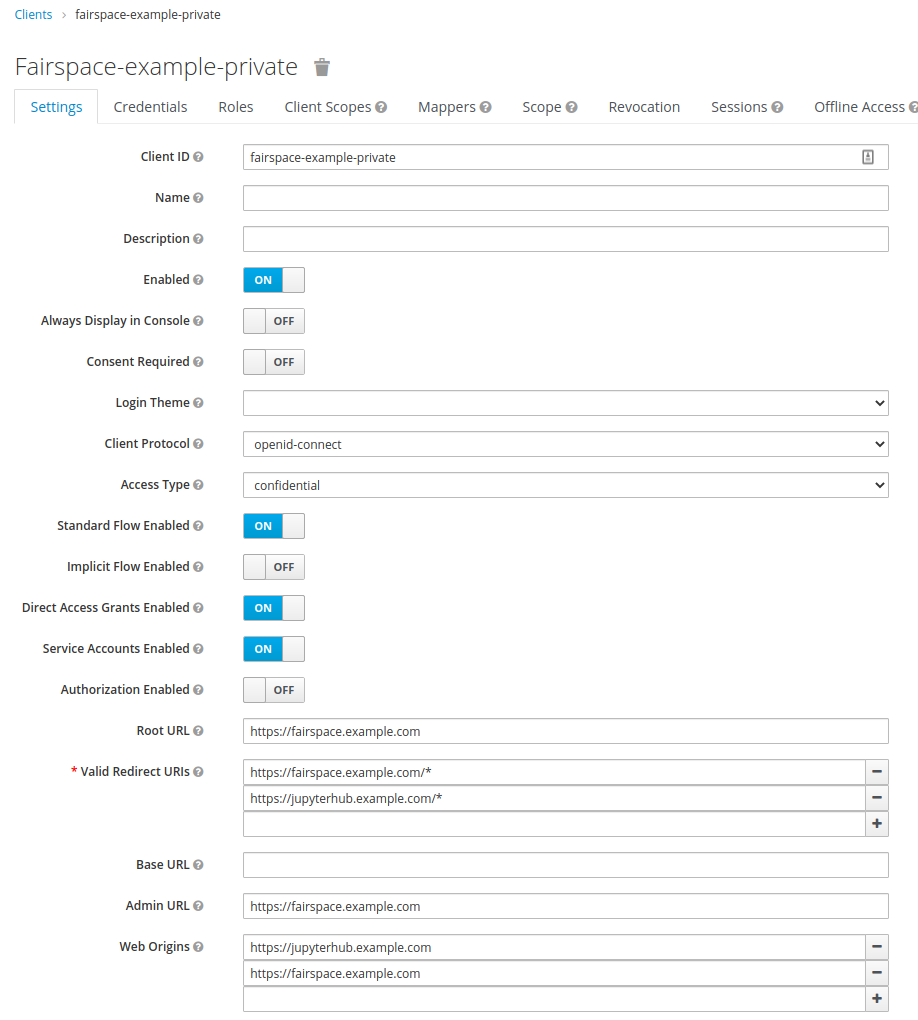
-
Set Access Type to confidential;
-
Set Service Accounts Enabled to On;
-
Ensure that
https://fairspace.example.comis added to the Valid Redirect URIs and Web Origins; -
Optionally (if you intend to add Jupyter Hub), ensure that the Jupyter Hub domain is added as well.
-
-
Assign the view-users role for client realm-management to the client service account:
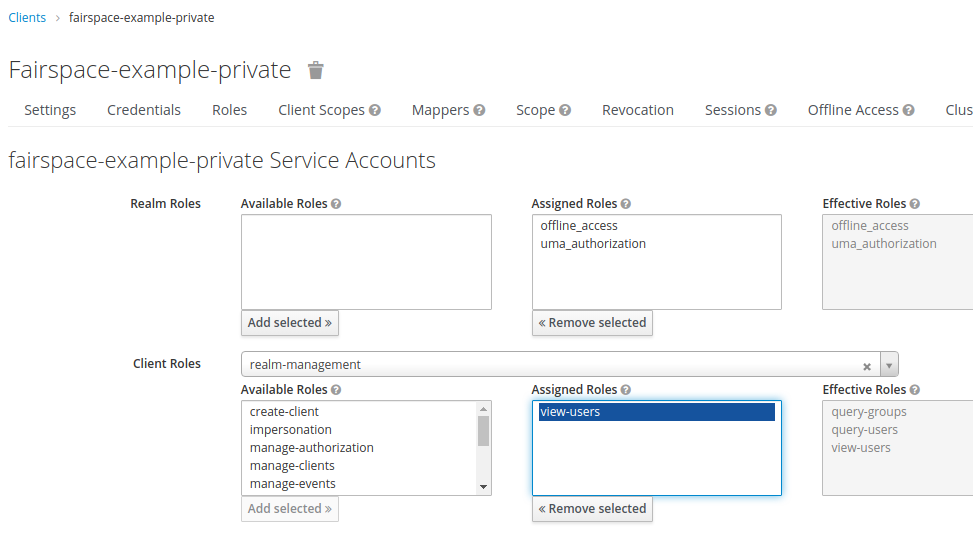
-
Copy the client secret from the Credentials tab, for use in the Fairspace configuration:
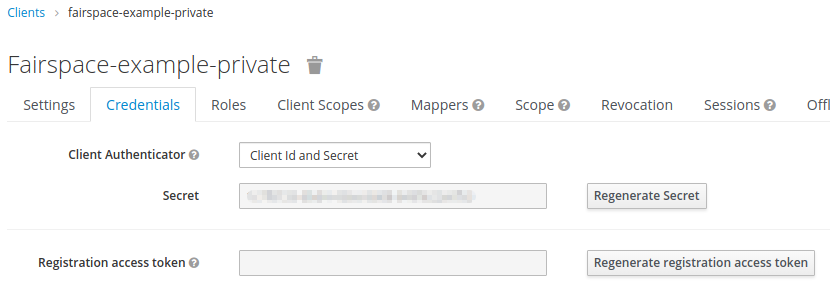
Deploy Fairspace
Create a new Kubernetes namespace:
kubectl create namespace fairspace-newCreate a new deployment (called release in helm terminology) and install the Fairspace chart:
~/bin/helm/helm install fairspace-new fairspace-2.0.2.tgz --namespace=fairspace-new \
-f /path/to/values.yaml --set-file saturn.vocabulary=/path/to/vocabulary.ttl --set-file saturn.views=/path/to/views.yamlYou can pass values files with -f and provide a file for a specified
value with --set-file.
Example values.yaml file:
# External dependencies for running the fairspace
external:
keycloak:
baseUrl: https://keycloak.example.com
realm: fairspace
clientId: fairspace-example-private
clientSecret: # Copy the client secret from Keycloak
# Settings for fairspace
fairspace:
name: "Example Fairspace"
description: "Example Fairspace"
ingress:
domain: fairspace.example.com
features: []
icons:
jupyter: "/icons/jupyter.svg" # path to the icon svg file
extra-icon: "extra-icon.svg" # path to the custom svg file
services:
jupyterhub:
name: "JupyterHub"
url: https://jupyterhub.example.com/user/${username}/lab
icon-name: "jupyter"
storages:
external:
name: external
label: "External storage"
url: https://storage.example.com/api/webdav
search-url: https://storage.example.com/api/search/files
root-directory-iri: https://storage.example.com/api/webdav
# Specific settings for Saturn subchart
saturn:
persistence:
files: # stores transaction logs and files
size: 60Gi
storageClass: expandable
database: # stores RDF database
size: 60Gi
storageClass: expandable
audit: # stores the audit log
size: 10Gi
storageClass: expandable
resources:
limits:
cpu: 1
memory: 16Gi
requests:
cpu: 500m
memory: 512Mi
image:
pullPolicy: Always
customStorageClass:
create: true
name: expandable
type: pd-standard
provisioner: kubernetes.io/gce-pd
allowVolumeExpansion: true
# Specific settings for Pluto subchart
pluto:
image:
pullPolicy: Always
socketTimeoutMillis: 600000 # 10 minutes
connectTimeoutMillis: 2000
maxFileSize: 1GB # max total size of file(s) that can be uploaded
maxRequestSize: 1GB # max total size of the request (should be > maxFileSize)
backends:
storageRoutes:
storage-external-webdav:
path: /api/storages/external/webdav/**
url: ${pluto.storages.external.url}
storage-external-search:
path: /api/storages/external/search/files/**
url: ${pluto.storages.external.search-url}
# Settings for a volume with PostgreSQL database used by Fairspace to store data for the metadata views.
postgres:
persistance:
storage:
size: 60Gi
storageClass: expandableAdditionally, to include custom icons for fairspace.icons option, you need to pass paths to the icon svg files as svgicons.<iconname>=<path/to/the/icon.svg using --set-file option:
~/bin/helm/helm install fairspace-new fairspace-2.0.2.tgz --namespace=fairspace-new \
-f /path/to/values.yaml --set-file saturn.vocabulary=/path/to/vocabulary.ttl --set-file saturn.views=/path/to/views.yaml --set-file svgicons.extra-icon=/path/to/extra-icon.svgIt is possible to pass multiple icon files with the --set-file option. Each of the icons has to be included as child of the svgicons key.
Deploy Jupyter Hub
Create a new Kubernetes namespace:
kubectl create namespace jupyterhub-newCreate a new deployment (called release in helm terminology) and install the Jupyter Hub chart:
~/bin/helm/helm pull oci://ghcr.io/thehyve/fairspace/helm-charts/fairspace-jupyter --version 0.8.9
~/bin/helm/helm install jupyterhub-new fairspace-jupyter-0.8.9.tgz --namespace=jupyterhub-new \
-f /path/to/values.yamlYou can pass values files with -f.
Example values.yaml file:
ingress:
domain: jupyterhub.example.com
# Specific settings for JupyterHub subchart
jupyterhub:
hub:
extraEnv:
JUPYTERHUB_CRYPT_KEY: xxx # A random string, you can use 'openssl rand -hex 32'
config:
FairspaceOAuthenticator:
client_id: fairspace-example-private
client_secret: # Copy the client secret from Keycloak
authorize_url: https://keycloak.example.com/realms/fairspace/protocol/openid-connect/auth
token_url: https://keycloak.example.com/realms/fairspace/protocol/openid-connect/token
userdata_url: https://keycloak.example.com/realms/fairspace/protocol/openid-connect/userinfo
logout_redirect_url: https://keycloak.example.com/realms/fairspace/protocol/openid-connect/logout?redirect_uri=https://jupyterhub.example.com
image:
pullPolicy: Always
singleuser:
image:
pullPolicy: Always
extraEnv:
TARGET_URL: https://fairspace.example.com
# EXTERNAL_TARGETS: external # Comma-separated list of names of external storages configured in Fairspace
proxy:
secretToken: # Generate strong secretUpdate an existing deployment
To update a deployment using a new chart:
~/bin/helm/helm upgrade fairspace-new fairspace-2.0.2.tgzWith helm upgrade you can also pass new values files with -f
and pass files with --set-file as for helm install.
Upgrading to a new version of the application may fail with a message, saying:
Forbidden: updates to statefulset spec for fields other than 'replicas', 'template', and 'updateStrategy' are forbidden
To prevent that, you can remove the existing StatefulSet before the upgrade:
kubectl delete statefulset fairspace-new --namespace fairspace-new --cascade=orphanThis will remove the StatefulSet, but keep all pods and volumes intact.
After running helm upgrade --install the StatefulSet will be recreated and the running pod will be replaced with a pod running the new application version.
Keep existing volumes on update of StatefulSet
Any changes on the StatefulSet will require a clean deployment, with new volumes. If you want to change the StatefulSet while keeping your existing volumes follow the procedure as described below.
To preserve your volumes get the volume id’s (pv-name’s):
$ kubectl get pv -n <namespace>Now patch the volumes so they are not deleted automatically when the VolumeClaim is removed:
$ kubectl patch pv <pv-name> -p '{"spec":{"persistentVolumeReclaimPolicy":"Retain"}}'
$ kubectl get pv # now shows status 'released'Uninstall the statefulset and remove the VolumeClaims. In released status the volumes cannot bind to a new claim, therefore remove the claim reference:
$ kubectl edit pv <pv-name>In the editor, remove the 'claimref' section.
The final step is to add the previously gathered volume id’s to the statefulset configuration:
volumeClaimTemplates:
- metadata:
name: postgres
spec:
accessModes: [ "ReadWriteOnce" ]
resources:
requests:
{...}
volumeName: "<volume-id>"Now execute the deployment.
Clean up deployment
To clean up an environment or completely reinstall an environment, you can use helm uninstall or kubectl delete.
| Be careful, you may lose data! |
Remove the application, but preserve persistent volume claims:
~/bin/helm/helm uninstall --namespace fairspace-old fairspace-oldPurge everything in the namespace, including persistent volume claims:
kubectl delete ns fairspace-oldLogging
For several purposes, three types of logs are generated:
-
Application log: Informative messages about the system state and application errors. This enables system administrators to diagnose problems.
-
Audit log: Records all user actions that add, change or delete data and access to files. This enables system administrators to audit important changes and access to sensitive data.
-
Transaction log: Detailed log of all database changes. This enables the system to restore the database if it is corrupted.
Do not change or remove this log!
As default, application and audit logs are written to standard output.
Additionally, the audit log is also written to log files in data/audit. Location can be overwritten by setting the AUDIT_LOG_ROOT environment variable.
The log files are automatically ‘rolled over’: today’s records are in audit.log, previous records are stored in daily log files with file name pattern
data/audit/audit.yyyy-MM-dd.log, and are retained for 50 days.
If the audit log needs to be kept for a longer period, the log configuration can be replaced, or log files can be transferred elsewhere, e.g., using filebeat.
The audit log is encoded in a JSON format, that can be processed by, e.g., logstash.
The transaction log is stored in data/log by default.
Configuration
The default log configuration for application and audit logs is in log4j2.properties.
The default can be overridden by placing a file log4j2.properties in the working directory where the application is run.
Audit log
The audit log is generated using log4j 2 and Mapped Diagnostic Context (MDC).
The basic idea of Mapped Diagnostic Context is to provide a way to enrich log messages with pieces of information that could be not available in the scope where the logging actually occurs, but that can be indeed useful to better track the execution of the program. Basically it groups log data from a single event (MDC.put) into a one log message.
Audit log entries contain several fields, including
event, user_name, user_email, user_id and request specific additional parameters.
Below we list the actions that are logged and which information is captured in the audit log.
WebDAV
Request: /api/webdav
| Request method | Event | Additional params | Description |
|---|---|---|---|
|
|
|
Read file |
|
|
|
Not used by the UI (metadata endpoint used instead) |
|
|
|
Create a new collection or directory. |
|
|
|
Copy a file or directory (executed on “paste” action) |
|
|
|
Rename a resource, move to a different location |
|
|
|
Mark a resource as deleted or delete permanently |
|
|
|
Not used by the UI (metadata endpoint used instead) |
|
|
|
Upload a file(s) or folder, undelete a resource, change collection status, access mode, owner or permissions. |
Events that are not logged: accessing collections, listing collection and directory contents, listing previous versions.
Metadata
Request: /api/metadata/
| Request method | Event | Additional params | Description |
|---|---|---|---|
|
|
|
Add all the statements in the given model to the database |
(Soft) |
|
|
Mark an entity as deleted |
|
|
|
Delete metadata |
|
|
|
Overwrite metadata in the database |
Fetching metadata is not included in audit log.
Workspace operations
Request: /api/workspace/
| Request method | Event | Additional params | Description |
|---|---|---|---|
|
|
|
Create workspace |
|
|
|
Delete workspace |
|
|
|
Add user to the workspace, change workspace user role, delete a user from a workspace |
Listing workspaces and workspace users is not included in the audit log.
User roles operations
Request: /api/users/
| Request method | Event | Additional params | Description |
|---|---|---|---|
|
|
|
Change organisation user role |
Fetching user information is not included in the audit log.
Data recovery using transaction log
In case of corruption the RDF database can be restored at any point of time using the transaction log.
Data recovery starts automatically on the Saturn application start, if the RDF database is empty and the transaction log containing entries is detected.
If Fairspace is deployed using Kubernetes, follow the steps below in order to restore the RDF database.
Stop (scale down) the application:
kubectl scale --replicas=0 --namespace fairspace-new statefulset/fairspace-newDelete a persistent volume consisting transient data.
If the reclaim policy of the pv is set to Delete or Recycle, delete the bound
persistent volume claim and the volume will be removed as well:
kubectl --namespace fairspace-new delete pvc database-fairspace-new-0If the reclaim policy is set to Retain, you need to delete the pv manually.
Find a name of pv bound to the deleted pvc (claim equals to database-fairspace-new-0):
kubectl --namespace fairspace-new get pv
and delete the pv:
kubectl delete pv <pv_name>
See the reclaim policy documentation of Kubernetes for more information.
Start (scale up) the application. The deleted pvc and pv will be recreated and data recovery will start automatically:
kubectl scale --replicas=1 --namespace fairspace-new statefulset/fairspace-newCheck the Saturn container’s logs to monitor the recovery process:
kubectl --namespace fairspace-new logs statefulset/fairspace-new fairspace-new-saturn -fThe logs should contain the following information:
2021-08-10 09:09:59 WARN Your metadata database is gone. Restoring from the transaction log containing 809 transactions
2021-08-10 09:09:59 INFO Progress: 0%
2021-08-10 09:10:20 INFO Progress: 1%
...
2021-08-10 09:27:17 INFO Progress: 99%
2021-08-10 09:27:17 INFO Progress: 100%
2021-08-10 09:27:17 INFO Committing changes
2021-08-10 09:32:36 WARN Restore is finished.
...
2021-08-10 09:32:39 INFO Saturn has startedAs soon as you see "Saturn has started" the RDF database should be restored to the state preceding the crash.
Architecture
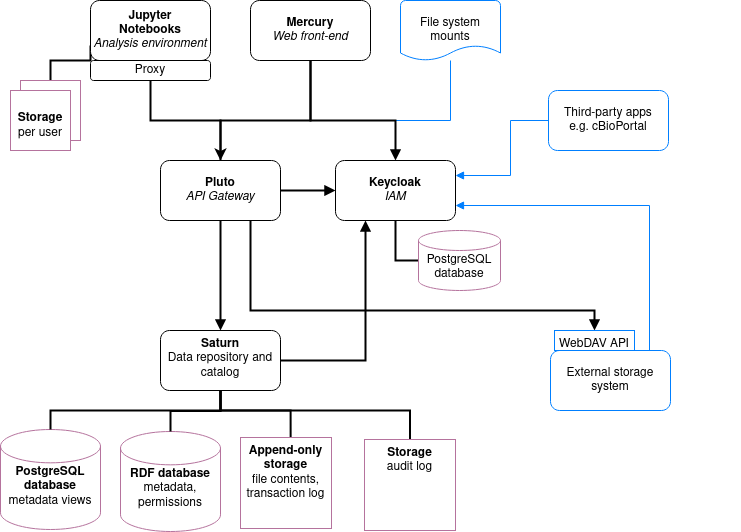
There are three core components of Fairspace: Mercury front-end, Pluto proxy and Saturn back-end. Together with storages and Keycloak as an identity and access management solution they create a complete Fairspace environment.
There could be additional applications integrated within the Fairspace environment, like Jupyter Hub and external storage systems.
Mercury
Web application developed using React and Material-UI component library. Mercury’s generic components for data and metadata allow changing the data model without a need to make changes in the front-end code.
Mercury connects to Pluto using session based authentication.
Pluto
Lightweight API gateway with OpenID Connect authentication support, that functions as a proxy between the microservices and the frontend. Pluto keeps track of user sessions, handles the OIDC authentication flow (authorization code based), JWT tokens (validation, refreshing) and redirects to back-end applications.
Pluto is a Java Spring Boot application, using Spring Cloud Gateway to route APIs and, together with Nimbus JOSE + JWT library, to connect to Keycloak, follow the authentication flow, handle web sessions and JWT tokens.
Saturn
The heart of Fairspace, back-end application handling all the logic and connection to the storages. Data repository and catalog.
Saturn is served as Apache Jena Fuseki SPARQL server, embedded into Jetty Web server.
Saturn uses Apache Jena RDF for handling RDF graphs, serialisation of triples and TDB triple store connection. All data within Fairspace, except to uploaded files and logs, are stored in Apache Jena RDF database.
For security, Saturn is integrated with Keycloak using Keycloak Jetty client adapter. It enables user authentication with access tokens.
Saturn is also responsible for authorization of users in Fairspace - it includes user access validators and storage of user records and permissions in the RDF database.
Saturn uses Milton IO, Java Webdav Server Library, to enable file storage management, directory listings and file properties handling, adding, moving, replacing and deleting files, directories and collections.
Storages
There are two types of storages that Saturn connects to: primary for storing RDF triples and file blobs, and secondary for efficient data retrieval.
There is also a possibility to configure an extra file storage for Saturn, separate from the primary one to keep e.g. search result exports created by users, so they could be mounted in the JupyterHub environment for further analysis.
Primary data storage
RDF database (Apache Jena TDB2) stores:
-
File metadata
-
Metadata entities
-
Permissions
-
User metadata
Data model schema is defined as a set of RDF classes and properties and SHACL constraints over them.
The virtual file system stores directory structure and file information in the RDF database and the files themselves in a separate append-only storage as blobs. Both adding a new file and updating an existing file result in adding a new file on the underlying storage. Every file version is stored as a separate blob and is normally never deleted (a deleted file is simply detached from the directory structure but remains in the blob store).
In combination with the fact that all operations on the RDF database are stored in the transaction log, that makes it possible to (incrementally) backup full state of workspaces and metadata at any point in time without stopping it.
Secondary data storage for efficient data retrieval
To improve the performance of metadata views, especially for counts and pagination, a simple relational database (PostgreSQL) is used. This database is flexible, created based on views configuration file and RDF storage, which enables recreation of the database at any point of time.
For models containing multiple attributes of Set/TermSet types, additional tables are created to store the values of these attributes. These tables are then left-joined with the main table to retrieve their values. However, if there are many such attributes, the performance of data retrieval from the views may degrade. To address this issue, materialized views are used for enhanced performance.
There are two types of materialized views: one for denormalized data, which includes the view ID and attribute data of Set/TermSet types, and another for joined views. For each joined view, there is one corresponding join materialized view (as specified in the views.yaml config).
Materialized views are refreshed during database reindexing, on Saturn initialization stage and metadata updates, provided that the doViewsUpdate flag is set to true in the metadata endpoints. The refresh is performed concurrently what allows for the system to be available during the update providing the old version of data until the new one is ready. To skip materialized views refresh on Saturn initialization stage, update the Saturn ConfigMap setting false value to viewDatabase.mvRefreshOnStartRequired.
Extra file storage
Similar to the primary file storage, the extra storage is a virtual file system that stores directory structure and basic file information in the RDF database. Unlike the primary one, the additional storage cannot be annotated with metadata from the defined model. Files themselves are stored in an extra storage as blobs. This storage however is NOT append-only. File blobs will be removed on deletion.
There is a basic, file-level access control over the storage - each user has "modify" access only to the files added by that user. Users cannot overwrite, read or even list each others files. This includes users with admin role.
To configure the extra storage to allow storage of metadata search results and to include the "Export to analysis" button in the user interface,
add ExtraStorage feature to the configured features list.
This will enable /api/extra-storage webdav endpoint with default analysis-export root directory created to store exported files.
Deployment architecture
Below you can find a diagram presenting the architecture of Fairspace deployment on a Kubernetes cluster, using Fairspace Helm chart described in previous sections.
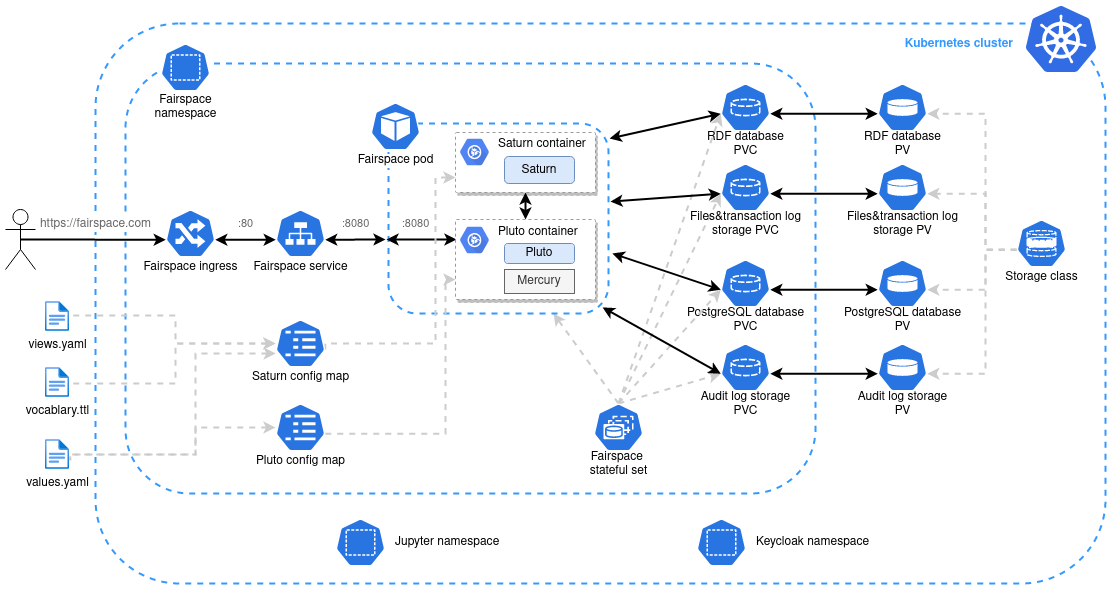
If Keycloak and JupyterHub are deployed together with Fairspace, they should be placed in separate namespaces of the same Kubernetes cluster.
For details on JupyterHub and Kubernetes see The JupyterHub Architecture documentation.
Acknowledgement
This project was realized thanks to and funded by:
-
The generosity of donors supporting the Curie Foundation.
-
Food Nutrition Security Cloud (FNS-Cloud). FNS-Cloud has received funding from the European Union’s Horizon 2020 Research and Innovation programme (H2020-EU.3.2.2.3. – A sustainable and competitive agri-food industry) under Grant Agreement No. 863059.
-
The Hyve BV, enabling open science (www.thehyve.nl).
License
Copyright (c) 2021 The Hyve B.V.
This program is free software: you can redistribute it and/or modify it under the terms of the Apache 2.0 License published by the Apache Software Foundation, either version 2.0 of the License, or (at your option) any later version.
This program is distributed in the hope that it will be useful, but WITHOUT ANY WARRANTY; without even the implied warranty of MERCHANTABILITY or FITNESS FOR A PARTICULAR PURPOSE. See the Apache 2.0 License for more details.
You should have received a copy of the Apache 2.0 License along with this program (see LICENSE). If not, see https://www.apache.org/licenses/LICENSE-2.0.txt.